Dr Bert Lampson – Bacteria: Deadly but Useful
Bacteria are found everywhere – in humans, animals and the environment – and are best known for being able to cause painful and even fatal infections. It may come as surprise, therefore, to learn that bacteria can also have useful applications. The lifetime work of Dr Bert Lampson at East Tennessee State University in the USA has focussed on the dangers and applications of bacteria. Over his career, Dr Lampson has discovered new mechanisms of antibiotic resistance, novel antibiotics, and bacterial proteins, with important applications for science and healthcare.
Revealing Antibiotic Resistance Mechanisms
Dr Bert Lampson began his research career as a graduate student in the early 1980s while studying at the University of Missouri-Columbia School of Medicine. At this time, scientists were developing new research technologies, including DNA sequencing. This process involves determining the sequence of nucleotides, the building blocks of DNA. The sequences can tell us what genes bacteria carry. Genes are inherited by bacteria and have many functions, such as encoding proteins that are used within the bacterial cell.
Dr Lampson used DNA sequencing to discover antibiotic resistance genes that are carried by plasmids. Antibiotic resistance genes are genes that encode mechanisms that lead to bacteria becoming resistant to antibiotics. Furthermore, plasmids are small circles of DNA that are physically separate from chromosomal DNA. The chromosomal DNA contains a large number of genes required to maintain cellular functions. In contrast, plasmids can carry a small number of additional genes, also known as accessory genes, that can help bacteria to adapt and survive in their environments. Furthermore, plasmids can be transferred amongst bacteria populations and spread genes. This has led to the spread of antimicrobial resistance genes that prevent antibiotic drugs from successfully treating an infection.
‘Wheel’ plate containing different Rhocococcus bacteria. Credit Bert Lampson.
The early work of Dr Lampson first involved sequencing a previously discovered plasmid in Staphylococcus epidermis. This bacterium is found around naturally on the skin. However, when the skin is broken the bacteria can enter the body and cause harmful infections. This particularly occurs when medical equipment, such as needles, are inserted into the body. These infections are most often treated with antibiotics.
Dr Lampson, under the guidance of Dr Joseph Parisi, showed that plasmids can induce resistance to three antibiotics (macrolides, lincosamide and streptogramin). This was demonstrated using transduction (gene transfer between bacteria using a bacterial specific virus called a phage), whereby the plasmids carrying the resistance genes were input into the bacteria not carrying the plasmid. Dr Lampson and his team then treated two sets of bacteria, those with and those without the plasmid, with antibiotics to see if carrying the plasmid led to bacteria becoming resistant.
Using DNA sequencing Dr Lampson and his fellow researchers pinpointed resistance to be due to a new gene, ermM. This gene has a similar sequence to a previously described resistance gene. However, the team observed changes upstream of the gene sequence, leading to changes in how the gene is expressed. ermM is constantly ‘switched on’, meaning it is continually expressed. The gene encodes an enzyme, a protein that is able to help the cell to carry out reactions. Critically, the protein alters the binding site, that is, the target site for antibiotics and prevents them from working.
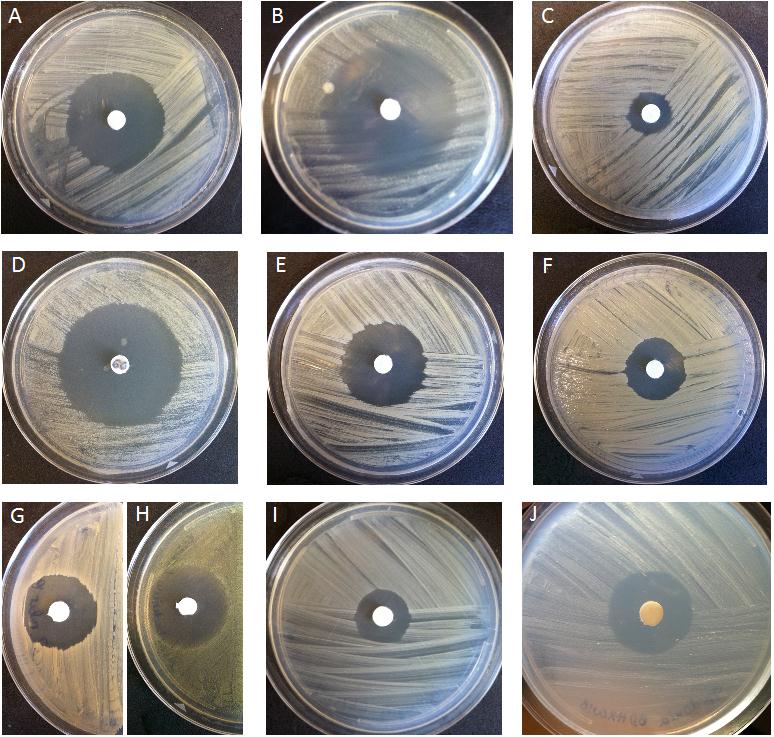

Spectrum of activity of the novel antibiotic from the Rhodococcus strain. Credit R. Borisova.
Remarkably, Dr Lampson also discovered a novel resistance mechanism that is carried by plasmids in the same species of bacteria. The newly discovered mechanism led to resistance to erythromycin. This is an antibiotic that is used to treat a number of infections. It operates by preventing bacterial cells from producing proteins that are essential for life, as well as replication.
Dr Lampson and his colleagues identified a new resistance gene that they designated erpA that was carried on a plasmid pNE24. Experiments that can detect specific DNA sequences, known as southern blot hybridisation, showed that the plasmid carries a new resistance mechanism. This is because the plasmid contains different DNA sequences to other plasmids that also carry erythromycin resistance genes. The new gene makes the bacterial cell less permeable, preventing the uptake of the antibiotic. This was shown by labelling the antibiotic with a radioactive carbon (Carbon-14) which helps scientists to detect the amount of antibiotic that the bacterial cells uptake.
This research holds important implications for understanding how bacteria become resistant to antibiotics and how this can spread amongst bacterial populations, particularly in hospitals.

Searching for Novel Antibiotics
When Dr Lampson started his role as a Research Scientist at East Tennessee State University (ETSU), he wanted to work on antibiotics again. However, this time, he took inspiration from work he had completed with the energy company, Energy Biosystems. This company was using an interesting soil bacterium, Rhodococcus, to try and produce cleaner fuels but Dr Lampson saw an additional opportunity for this research. Although ETSU is not a primarily research university, Dr Lampson was also keen for students to participate in this exciting new research project.
This work focused on exploiting Rhodococcus for antibiotics, which at first, may seem paradoxical. Why would bacteria produce antibiotics that are used to kill them? The reason is that species of soil bacteria have to compete with one another to survive. Therefore, bacteria produce antibiotics to outcompete other species.
Dr Lampson and his students grew Rhodococcus on plates that contain agar, on which bacteria can survive. They extracted molecules from these agar plates and then they searched for molecules and proteins from Rhodococcus using different forms of chromatography. This process helps to separate the different molecules from one another. After separating the extracts, the team coated small disks with the extracts and tested whether the bacteria could survive in its presence.
This work led to the discovery of a new compound that could inhibit the growth of bacteria with potentially significant applications in medical and veterinary practice. As such, this key study resulted in several more research projects, and towards the end of 2018, a publication for Dr Lampson and his students in the high impact journal PLoS ONE.

Discovering the First Bacterial Reverse Transcriptase
Dr Lampson has additional research interests outside the field of antibiotic resistance. Notably, whilst working as a postdoctoral researcher at the Robert Wood Johnson Medical School, he discovered a protein with very important applications.
Using soil bacteria, this time Myxococcus, Dr Lampson discovered the first example of a bacterial reverse transcriptase. This is a protein, also known as an enzyme, that is responsible for forming DNA from RNA. Generally, people associate DNA with producing RNA and RNA with making proteins. However, this is a simplified view – in reality, the process is much more complex.
Reverse transcription is the formation of DNA from RNA and this takes place in most organisms, including humans and bacteria. Interestingly, it is believed that reverse transcription was a key process in the origins of life and it is hypothesised that RNA was the source of genetic information and cellular functions. This collectively is known as the ‘RNA world’.
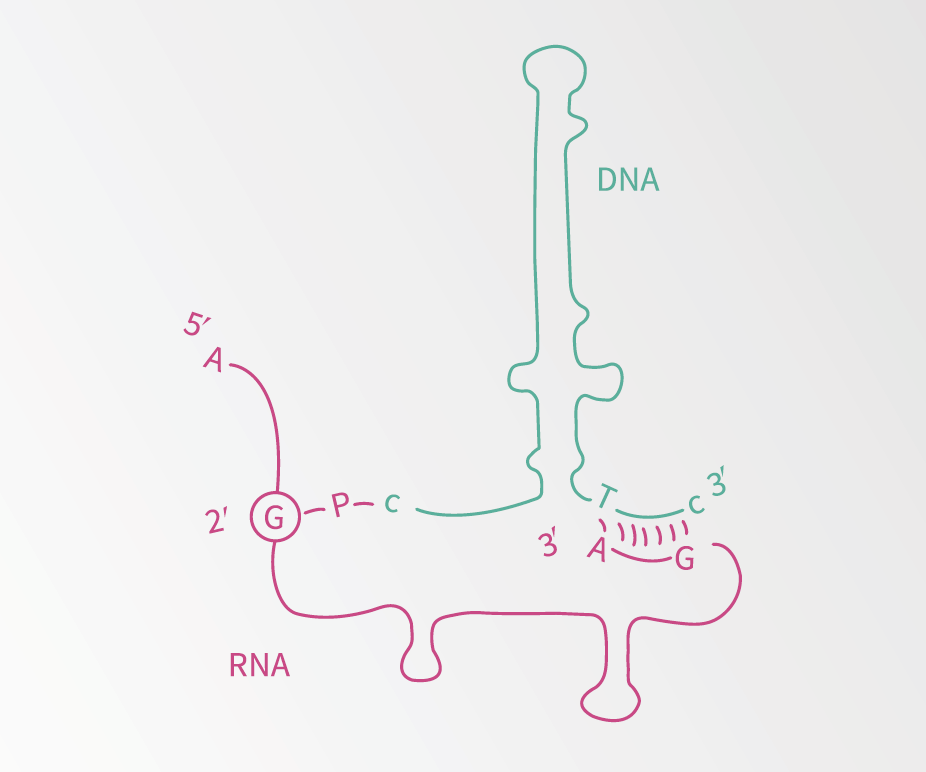
Illustration of msDNA.
Dr Lampson, under the mentorship of Professor Masayori Inouye, discovered bacterial reverse transcriptase after the discovery of another strange bacterial element, msDNA. This molecule contains both RNA and DNA bonded together. Dr Lampson and his colleagues showed that this molecule was produced by reverse transcriptase, making this the first reverse transcriptase to be identified in bacteria. Importantly, reverse transcriptase can be used to make DNA from RNA, which in turn means the DNA can be identified using a process known as polymerase chain reaction (PCR) or DNA sequencing. This allows scientists to figure out what DNA is being expressed.
These findings further led to the discovery of a thermostable reverse transcriptase that can operate at high temperatures. High temperatures otherwise alter the structure of a protein and prevent it from functioning. Although reverse transcriptase had been identified from viruses previously, heat stable bacterial reverse transcriptases are more advantageous because they increase the accuracy of DNA synthesis and are potentially more stable under laboratory experimental conditions.
Dr Lampson and his team searched for the genome sequence of Bacillus stearothermophilus. This bacterium is thermophilic, meaning it can survive at extreme temperatures. They encountered a gene that is similar to genes encoding group II introns that have reverse transcriptase activity, and also provided supportive evidence for the ‘RNA world’ hypothesis. After the scientists cloned the gene and input into Escherichia coli to produce the protein, they confirmed it has reverse transcriptase activity and could operate at temperatures as high as 75oC. Dr Lampson and ETSU have now patented the reverse transcriptase and have licensed it to a company called InGex that developed it as a commercially available enzyme called TGIRT III.

Dr Lampson continues his research and lecturing at ETSU. His current research largely focusses upon looking at the metabolism of soil bacterium Rhodococcus and potential antibiotics produced by this bacterium. He also continues to take interest in reverse transcriptase and their diversity amongst bacteria. Collectively, his work shows that although bacteria can cause harmful, resistant infections, they also harbour proteins with useful applications that can be exploited and bring important benefits.
Reference
https://doi.org/10.33548/SCIENTIA630
Meet the researcher

Dr Bert Lampson
Department of Health Sciences
East Tennessee State University
Johnson City
Tennessee, TN
USA
Dr Bert Lampson received his PhD in Medical Microbiology from the University of Missouri-Columbia School of Medicine in 1986. He then undertook postdoctoral research in the laboratory of Dr Masayori Inouye at the University of Medicine and Dentistry, Robert Wood Johnson Medical School. In 1998, Dr Lampson became an Assistant Professor in the Department of Health Sciences at East Tennessee State University which was followed by promotion to his current role of Associate Professor in 2004. He has had a successful research career spanning four decades, during which time he has contributed significantly to tackling the worldwide problem that is antibiotic resistance.
CONTACT
E: lampson@etsu.edu
KEY COLLABORATORS
Dr Masayori Inouye, Robert Wood Johnson Medical School
Dr Abbas Shilabin, East Tennessee State University
Professor Joseph T. Parisi (deceased)
Graduate students: J. Vellore, S. Moretz, T. Barber, M. Carr, R. Borisova, A. Ward and K. Sellick
FUNDING
Takara Bio
East Tennessee State University Research Foundation
Department of Health Sciences, ETSU
FURTHER READING
AL Ward, P Reddyvari, R Borisova, AG Shilabin, BC Lampson, An inhibitory compound produced by a soil isolate of Rhodococcus has strong activity against the veterinary pathogen R. equi, PLoS ONE, 2018, 13(12), e0209275.
BC Lampson, M Inouye, S Inouye, Retrons, msDNA and the bacterial genome, Cytogenetic and Genome Research, 2005, 110, 491–499.
J Vellore, SE Moretz, BC Lampson, A group II intron-type open reading frame from the thermophile Bacillus (Geobacillus) stearothermophilus encodes a heat-stable reverse transcriptase, Applied and Environmental Microbiology, 2004, 70, 7140–7147.
JT Parisi, J Robbins, BC Lampson, HW Hecht, Characterization of a macrolide lincosamide, and streptogramin resistance plasmid in Staphylococcus epidermidis, Journal of Bacteriology, 1989, 148, 559–564.
BC Lampson, JT Parisi, Naturally occurring Staphylococcus epidermidis plasmid expressing constitutive macrolide-lincosamide-streptogramin B resistance contains a deleted attenuator, Journal of Bacteriology, 1986, 166, 479–483.
BC Lampson, JT Parisi, Nucleotide sequence of the constitutive macrolide-lincosamide-streptogramin B resistance plasmid pEN131 from Staphylococcus epidermidis and homologies with Staphylococcus aureus plasmids pE194 and pSN2, Journal of Bacteriology, 1986, 167, 888–892.
BC Lampson, W von David, JT Parisi, A novel mechanism for plasmid-mediated erythromycin resistance is conferred by pNE24 from Staphylococcus epidermidis, Antimicrobial Agents and Chemotherapy, 1986, 30, 653–658.

Want to republish our articles?
We encourage all formats of sharing and republishing of our articles. Whether you want to host on your website, publication or blog, we welcome this. Find out more
Creative Commons Licence
(CC BY 4.0)
This work is licensed under a Creative Commons Attribution 4.0 International License. 
What does this mean?
Share: You can copy and redistribute the material in any medium or format
Adapt: You can change, and build upon the material for any purpose, even commercially.
Credit: You must give appropriate credit, provide a link to the license, and indicate if changes were made.
More articles you may like
How Food Environments Shape Our Eating Habits
How we eat dramatically impacts our health, yet millions of Americans live in ‘food deserts’ – areas with limited access to fresh, nutritious food. Recent research reveals that solving this crisis requires looking beyond just physical access to food to understand how our entire community environment shapes our dietary choices. Through a series of pioneering studies, Dr Terrence Thomas and colleagues at North Carolina A&T State University have been investigating how different aspects of our food environment influence what we put on our plates. Their findings suggest that creating lasting change requires reimagining how communities engage with food at every level.
Probing Electron Dynamics in the Ultrafast Regime
In the atoms that make up the matter around us, negatively charged particles called electrons have properties such as spin and orbital angular momentum. Researchers at Martin Luther University Halle-Wittenberg have developed a theoretical framework which allows them to simulate the dynamics of the spin and orbital angular momentum of electrons in materials when probed with an ultrafast laser pulse. Using this framework, they are able to simulate different materials and improve our understanding of dynamics on an atomic scale.
Seeing Beneath the Surface: Exploring Deltaic Reservoirs with Augmented Reality
In the Aínsa Basin of the Spanish Pyrenees, the Mondot-1 well was drilled, cored, and fully logged to capture a detailed record of a long-buried ancient river delta system. Dr. John D. Marshall, Dr. Jürgen Grötsch, and Dr. Michael C. Pöppelreiter with co-workers at Shell International used this core to trace how sediments once flowed across the landscape, and were deposited under shifting tectonic conditions. The team employed augmented reality and interactive virtual displays; these innovative tools offer new ways to explore subsurface depositional systems, and are particularly useful in locations where physical access to the core is difficult, or no longer possible.
Dr Jim Wu | Ziresovir Offers New Hope for Treating Respiratory Syncytial Virus Infections
Respiratory syncytial virus (RSV) causes respiratory tract infections in children and adults. While for many patients the outcomes of infection are mild, for others, infection can prove fatal, and there is a lack of effective treatments. Dr Jim Wu from the Shanghai Ark Biopharmaceutical Company in China carries out his vital research to develop new, safe, and effective treatments to tackle this killer.




