Dr Shigeki Iwase – Neurodevelopmental Disorders Arising from Histone Methylation Malfunctions
Neurodevelopmental disorders range from those on the relatively common autism spectrum to much rarer disorders such as KDM5C-disorder and Weidemann-Steiner Syndrome. Exciting advancements in human genetics have shown that histones – the proteins our DNA wraps around – play a vital role in healthy brain development. Dr Shigeki Iwase from the University of Michigan studies how mutations in the enzymes that regulate histone structure and function can cause cognitive disorders. His work has led to important new discoveries, including how counterpart enzymes can be utilised for therapies.
Histones Condense the Genome
If you stretched out all the DNA contained in just one of your cells, it would create a thread around two metres long. However, safely within the nucleus of your cells, your DNA is coiled up tightly into the recognisable X shape of the chromosomes. To keep genetic material condensed and stable, it wraps around proteins called histones approximately 1.7 times. These proteins are found in groups of eight; when DNA has wrapped round one of these groups, it is known as a nucleosome. Many nucleosomes in repeating subunits create chromatin. As a result of all this wrapping and condensing, your genetic material in one cell, in the form of chromatin, is only around nine centimetres long.
The group of eight histones has a specific configuration and is made up of five variations of the protein – H1, H2A, H2B, H3 and H4. Because each histone protein is positively charged, they each bind strongly to DNA strands, which are negatively charged, thanks to the phosphate group in their backbone.
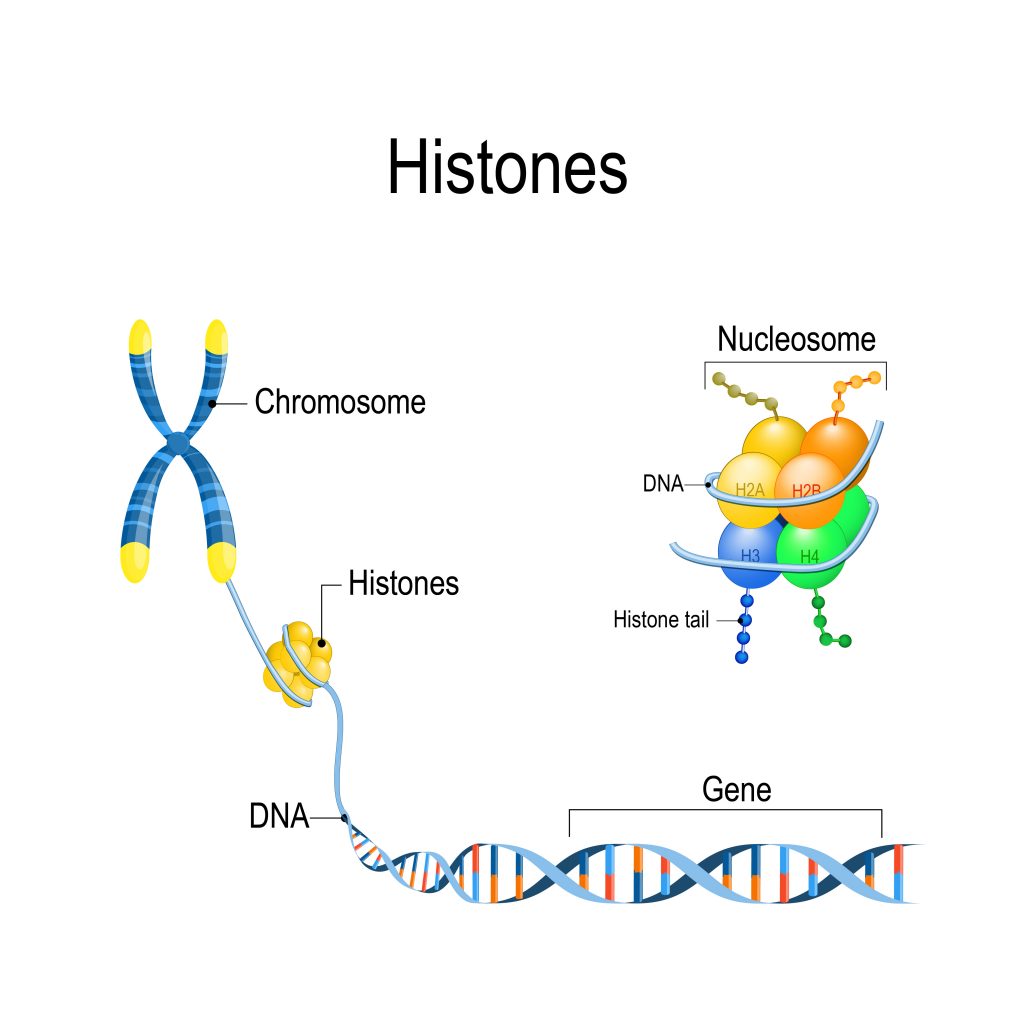
Histones
Modifying Histones for Cell Regulation
Histones also have an important role in regulating gene expression, which means controlling when and which parts of the genome are permitted to be transformed into proteins. When necessary, the proteins go through a process called covalent post-translational modification which alters the structure of chromatin, therefore impacting gene expression. Examples of post-translational modifications include phosphorylation, acetylation, ubiquitylation and methylation, all of which involve attaching a new molecule, compound or protein to the histone tail.
During methylation, enzymes called histone methylases attach one, two or three methyl groups to amino acids on the H3 and H4 proteins. It is therefore called mono-, di- or trimethylation, accordingly. These methyl groups are added to a lysine amino acid, which is abbreviated to ‘K’ or an arginine amino acid, abbreviated to ‘R’. Depending on how many methyl groups are attached and to which amino acid, the resulting nucleosome structure can be weakened. If the attractions between the DNA strand and the histone are diminished, the DNA will start to uncoil from the nucleosome which makes it much easier for RNA polymerases (transcription enzymes) to join on and begin the process of transcription. In this way, histone methylation can turn genes ‘on’. On the other hand, methylation of a lysine at the 9th position on H3 is a transcriptional silencing signal, and that gene will be turned ‘off’ for the time being.
The important enzymes which add on histone methylation are known as writer enzymes and their counterparts are eraser enzymes, which remove them. As with any other enzyme in our bodies, enzymes are proteins coded for by our genetics and are, therefore, subject to mutations that can alter our physiology. Studying this phenomenon, and how it may lead to intellectual disabilities and autism spectrum disorders, is Dr Shigeki Iwase from the University of Michigan in Ann Arbor. His research into how histone methylases and demethylases are vital for cognitive function has uncovered fascinating new information in his field.

Discovering a Gene Connected to X-linked Diseases
As the world of genetics research broadens and the knowledge of our inner workings deepens, scientists have realised that the histones within our nuclei can play an important role in brain development. A well-known modification to histones is the methylation of lysine 4 on H3 (H3K4me), which promotes transcription.
Initial work by Dr Iwase and his team led to the discovery of a new gene labelled KDM5C. This gene codes for a H3K4me2/3 demethylase, which is an enzyme that removes methyl groups from H3K4 when it is already di- or tri-methylated. Removing the methyl groups that would promote transcription means that this enzyme is a transcriptional repressor – it prevents transcription. The team found that when KDM5C is mutated in humans, it leads to a disorder called mental retardation, X-linked, syndromic, Claes-Jensen type. This condition is characterised by significantly diminished intellectual functioning, seizures, aggression and reduced height, all presenting during brain development.
Now called X-linked intellectual disabilities (instead of X-linked mental retardation), this family of disorders is inherited recessively through the female X chromosome. Because females have two X chromosomes, they can either inherit two mutated alleles (versions of the gene) and therefore express the disorder, or inherit one mutated allele and become just carriers of the disease. If only one of their X chromosomes carries the mutation, the ‘healthy’ one usually nullifies the diseased one. However, males hold only one X chromosome inherited from their mother and if they receive a mutated gene from her, will present with the X-linked disease. As a result, X-linked recessive diseases are more commonly found in males.
‘… the yin-yang action of writer-eraser enzymes can be targeted to ameliorate neurodevelopmental disorders.’
Mice and Humans with the Same Genetic Mutation Show the Same Symptoms
Building on this research, Dr Iwase investigated how mutations in KDM5C would affect mice and whether the consequences were comparable to humans with KDM5C mutations. The team knocked out certain sections within this gene (exons 11 and 12) to make it ineffective. Subsequently, the mice exhibited a smaller body size and diminished body weight. Interestingly, in humans, around 60% of individuals who are known to have a KDM5C mutation have a shorter than average stature.
Mice containing the mutation were significantly more aggressive than their littermates without it, were much less social and showed some memory impairment. All of these traits are found in many individuals with autism who hold KDM5C mutations and in those with X-linked intellectual disabilities.
Dr Iwase also found various neuropathway abnormalities in KDM5C knockout mice. Dendritic arborisation, or dendritic branching, is an essential part of brain development whereby neurons branch out to form new synapses. This was shown to be distorted in the mice, as they exhibited shorter and thinner dendrites in their basolateral amygdala – a centre in the brain for perceiving and regulating emotion, fear and aggression.
Overall, this study demonstrated how important KDM5C is for regulating pathways that lead to the normal development and function of neuronal circuitries. All of the data gathered from this experiment strongly suggest that mutations to KDM5C cause X-linked intellectual diseases because the loss of its function leads to gene expression that presents as clinical disease. The team believes that this is not just the case in mice but humans as well, as the cognitive deficits and behaviour patterns are so alike.

‘The Yin-yang Action of Writer-eraser Enzymes’
More recently, Dr Iwase examined whether a faulty histone enzyme could be modulated by using a different, functioning one.
The methylation processes of H3K4 are regulated by thirteen different enzymes but their functions in the brain are largely unknown. Genetic mutations in nine enzymes have strong associations with neurodevelopmental disorders, which are also known as brain H3K4 methylopathies in this context. Clinical genetics studies have told us that these enzymes are important in brain development.
Theoretically, as methylation modifications carried out by these enzymes are reversible, it could be possible to correct any errors by modulating the writer and eraser enzymes associated with them. Dr Iwase again used the eraser KDM5C enzyme which removes methylation, as well as including the writer enzyme KMT2A, which adds it. If a person is ‘haploinsufficient’ for the KMT2A gene, it means that they do not produce enough of the protein for proper function. Weidemann-Steiner Syndrome presents as a result, which causes intellectual disability and speech delay, short stature, characteristic facial features and hypotonia (decreased muscle tone).
Both of these enzymes should be found at high levels across the brain during development so that they can interact to regulate methylation. In a series of experiments on mice, the team found some fascinating results. In KDM5C-deficient mice, inhibiting KMT2A ameliorated many of the usual cognitive defects. So, the mice who could not properly remove methylation from histones when necessary, were prevented from increasing methylation which would otherwise result in disease. Likewise, cognitive issues arising from KMT2A mutation were alleviated by KDM5C deletion. It appears that controlling one of these partner enzymes amends the dysfunctions of the other, leading Dr Iwase to conclude, ‘…the yin-yang action of writer-eraser enzymes can be targeted to ameliorate neurodevelopmental disorders.’
Dr Iwase has helped the scientific community deepen its understanding of how methylation of histones – and dysfunctions in the process –can impact cognitive health and function. Importantly, his findings have exciting implications for the use of writer-eraser enzymes as potential remedies for neurodevelopmental disorders in the future.
Reference
https://doi.org/10.33548/SCIENTIA643
Meet the researcher

Dr Shigeki Iwase
Department of Human Genetics
University of Michigan, Medical School
Ann Arbor, MI
USA
Dr Shigeki Iwase completed his Bachelor of Science at the University of Tsukuba in Japan, before achieving his PhD from the same university in 2006. He then completed his postdoctoral training at Harvard Medical School in Boston in 2012. Since then, he has received multiple awards and published numerous papers on his studies. Dr Iwase now serves as an Associate Professor in Human Genetics at the University of Michigan in Ann Arbor, where he also carries out his research. His work focusses on chromatin in our genetic material and its involvement in health and disease in the brain.
CONTACT
E: siwase@umich.edu
W: https://medicine.umich.edu/dept/human-genetics/shigeki-iwase-phd
Twitter: @Iwase_Lab
KEY COLLABORATORS
Natalie Tronson (University of Michigan)
Jun Xu (Washington State University)
Yang Shi (Harvard Medical School)
FUNDING
National Institute of Neurological Disease & Stroke
Basil O’Connor Starter Scholar Research Awards from March of Dimes Foundation
Farrehi Family Foundation
FURTHER READING
CN Vallianatos, B Raines, RS Porter, et al., Mutually suppressive roles of KMT2A and KDM5C in behaviour, neuronal structure, and histone H3K4 methylation, Communications Biology, 2020, 3(1), 1–14, doi:10.1038/s42003-020-1001-6
S Iwase, E Brookes, S Agarwal, et al., A Mouse Model of X-linked Intellectual Disability Associated with Impaired Removal of Histone Methylation, Cell Reports, 2016, 14(5), 1000–1009, doi:10.1016/j.celrep.2015.12.091
S Iwase, F Lan, P Bayliss, et al., The X-Linked Mental Retardation Gene SMCX/JARID1C Defines a Family of Histone H3 Lysine 4 Demethylases, Cell, 2007, 128(6), 1077–1088, doi:10.1016/j.cell.2007.02.017

Want to republish our articles?
We encourage all formats of sharing and republishing of our articles. Whether you want to host on your website, publication or blog, we welcome this. Find out more
Creative Commons Licence
(CC BY 4.0)
This work is licensed under a Creative Commons Attribution 4.0 International License. 
What does this mean?
Share: You can copy and redistribute the material in any medium or format
Adapt: You can change, and build upon the material for any purpose, even commercially.
Credit: You must give appropriate credit, provide a link to the license, and indicate if changes were made.
More articles you may like
Prof Candis M. Morello – Prof Jan D. Hirsch | Recent innovations in pharmacy education
A pioneering research team from the Skaggs School of Pharmacy and Pharmaceutical Sciences, University of California, San Diego, United States, has been instrumental in developing innovative techniques for teaching pharmacy students. The Next Generation of Pharmacist Educators (NextGen-RxEd) programme is a new method of training the next generation of pharmacist educators and academics. To help pharmacists and pharmacy students visualise the complex issues experienced by their patients, the team led by Professors Candis Morello and Jan Hirsch developed an innovative educational tool, called the Medication Therapy Management (MTM) Spider Web.
Professor Michael Yarus | How RNA Started the Conversation That Built Life
The genetic code stores the instructions for building proteins, yet how it first arose remains unclear. It likely did not appear fully formed, but instead emerged step by step from simple chemical interactions. In this study, a team led by Professor Michael Yarus at the University of Colorado Boulder shows how early ribonucleic acid (RNA) molecules could bind specific amino acids, creating the first coding relationships. These early interactions are then refined through evolutionary processes such as duplication and merging of partial codes. By combining the experimental data with computer simulations, the work provides a testable pathway from prebiotic chemistry to the modern genetic code.
Identifying Nutritional Risk in Early Childhood: Insights from NutriSTEP®
Early childhood is a critical period for growth and development, yet many young children face nutritionrelated risks that can go unnoticed. Professor Janis Randall Simpson and colleagues have developed NutriSTEP®, validated and reliable screening tools that help identify potential nutritional concerns in toddlers and preschoolers. Their large-scale analysis of Canadian data reveals patterns in diet, behaviour, and food access that could help guide early interventions and support healthy development.
Professor Tian Yu Cao | Twistor Theory: A New Framework for Quantum Gravity
At Boston University, Professor Tian Yu Cao is rethinking the foundations of modern physics. His work builds on twistor theory which demonstrates that spacetime is secondarily derived from twistor constructions, but goes further to highlight the most important implication of the Penrose transform in that the primary physical agents can only be mathematically described by elements of cohomology with the defining feature having roots in spin. This view of primary agents combines with Cao’s other major claim that quantum behaviour itself may arise from the physical property of spin leads to a new consistent framework of quantum gravity in which long-standing puzzles in black holes (evaporations) and cosmology (transitions between cycles of cosmos) can be adequately addressed, with the crucial help from the on-going development of operator product expansion formular defined on twistor space.




