Dr Craig Albertson – Beyond Genetics
Shifting the Central Dogma
The classic Central Dogma of biology provides a pathway for how genetic information flows from DNA to yield a phenotype. This flow follows the transcription of DNA into RNA, which is subsequently translated into protein. Thus, as plainly stated, the Central Dogma recognises a gene-centric view of a biological system. This means that, based on the Central Dogma, all of the information necessary to define the state of an organism is contained in the sequence of its genome. This is actually a pretty fair assessment of molecular biology when one is describing simple single-cellular organisms, such as bacteria. However, this gene-centric view does not hold up when studying complex multicellular organisms. It is well known for example that monozygotic twins, with identical genomes, can act, behave and even look quite different. Clonal lines of plants grown under drought or wet conditions will display very different and stable phenotypes. Butterflies that metamorphose in spring will have more dark pigment on their wings compared to the same species that metamorphose in the summer. These observations suggest that there is further information that is responsible for the production and maintenance of phenotypes.
Given that the Central Dogma does not provide a complete explanation for biological processes in various fields of biology, many disciplines are shifting this paradigm to better describe biology. One such discipline is evo-devo – a field of biology that combines evolutionary biology and developmental biology. Many researchers in this field believe that the basic Central Dogma cannot define complex traits; for example, if gene expression was simply occurring as the Central Dogma describes, then scientists could reconstruct a human face from a DNA sample. This would mean that DNA from a crime scene could be used to generate a facial image of a suspect, the remains of prehistoric people could be used to reconstruct their faces or the face of a baby could be predicted before birth using DNA from the amniocentesis. This is an exciting idea, but in reality it’s actually quite farfetched. There are several reasons that DNA alone cannot predict facial shape.
‘How the antifreeze glycoproteins, as well as other antifreeze proteins, evolved into functional antifreeze molecules is of much interest to evolutionary biologists’
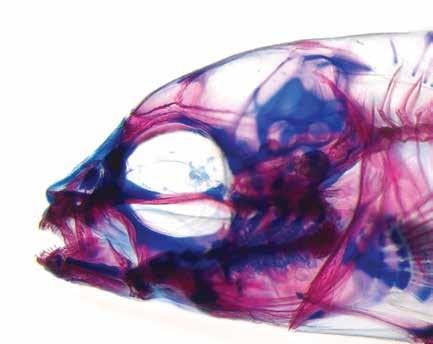

Variation in a single locus can alter multiple aspects of a dynamic system
Traditionally, the focus of evolutionary developmental biology was to link genes with straightforward shifts in morphology, but in order for a species to adapt to new environments, a divergence in resource use must also occur. This divergence often involves variation in multiple components of complex functional systems. Based on this idea, Dr Albertson and his colleagues predicted that, for evo-devo research, the elucidation of broad evolutionary processes should investigate more dynamic traits within this framework.
Dr Albertson’s work focuses on the diversification of craniofacial morphology and during vertebrate evolution. The adaptive variation of craniofacial structure occurs in response to different food sources and habitats, which in turn plays a role in niche partitioning and speciation. Actually, a majority of the morphological and functional divergence between vertebrates occurs in the craniofacial region. Thus, it’s no surprise that numerous studies have examined the patterns of craniofacial divergence in a wide variety of animals. Of the animals studied, East African cichlids are considered an excellent system to study the genetic and developmental mechanisms that promote craniofacial divergence, because they exhibit a large degree of variation in craniofacial morphology.
To study variations in the craniofacial skeleton, Dr Albertson and his team combined traditional quantitative trait locus mapping, population genetics and experimental embryology to understand how differences in gene expression during larval development in African cichlids contributes to adaptive morphological variation in adults. In one study, Dr Albertson’s team showed that a gene in the Hedgehog (Hh) signalling pathway mediates widespread variation in bones that affects the kinematics of lower jaw depression in cichlids. This seemingly simple action is controlled by a complex arrangement of bones and tendons, and the efficiency of this system has major implications for how well an animal can feed. This finding contributes to the one of the key questions in evolutionary studies, which asks how genetic variation translates into ecomorphological adaptation and ultimately, fitness. Dr Albertson’s study presents experimental data that variation in a single gene affects various aspects of a dynamic mechanical system. Ultimately, this work links adaptive variations at genetic, developmental and functional levels.
Epigenetics – a divergence from the Central Dogma
Epigenetics is the study of cellular and physiological phenotypic trait variations that are caused by external or environmental factors that turn genes on and off. While the study above highlights the genetic roles for adaptive variation in the jaw, these genetic effects only contribute to a relatively small percentage of the phenotypic variation that is observed. Cichlids rapidly evolve and display a large spectrum of variation in their craniofacial skeletons, and Dr Albertson is interested in how these changes occur at the genetic level. However, his team also found that there is an epigenetic origin for the adaptive variation that occurs in the cichlid jaw.
The team revealed that cichlid larvae begin gaping their mouths right after the cartilaginous lower jaw forms, which is just prior to the commencement of bone development. They showed further that the frequency of gaping varies between species in a manner that predicts variations in bone deposition. Moreover, when the gaping is disrupted in the fast gaping species, the jaw forms in a manner that is similar to the slow gaping species. In contrast, when the gaping is forced to occur at a higher frequency, the jaw of the slow gaping species develops to resemble that of the fast gaping species. Finally, Dr Albertson and his colleagues showed that these epigenetic changes also occur through the Hh signalling pathway. Thus, both the genetic and epigenetic path to jaw shape variation appears to leverage the same pathway. Taken together, these results emphasise the complexity of how craniofacial shape takes place and propose a novel experimental framework for investigating sources of phenotypic variation beyond those determined by changes in gene sequences.
‘While we know a lot about how the head develops, we know much less about how the complex geometry of the skull is determined’
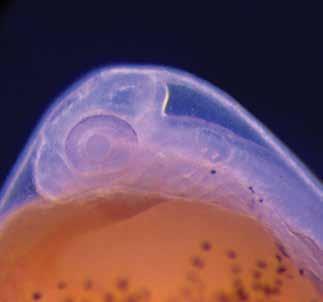
Environmental influences on gene deployment during trait development
Phenotypic plasticity refers to the ability of organisms to alter their phenotype in response to changes in the environment. Recently, phenotypic plasticity has been a hot topic in discussions of evolutionary potential, but a complete understanding of the genetic basis of plasticity is lacking. Dr Albertson and his team investigated the genetic basis of phenotypic plasticity in cichlid fishes. They crossed two divergent species to generate a genetic mapping population. These hybrid families, at early juvenile stages, were split and raised in alternate foraging environments that mimicked benthic/scraping or limnetic/sucking modes of feeding. The different environments produced variations in morphology that were largely comparable to the major axis of divergence among the cichlids, which supported the flexible stem theory of adaptive radiation – a theory proposing that patterns of plasticity in a population will shape future patterns of morphological evolution. In addition, the study revealed that the genetic architecture of nearly every morphological trait examined was highly sensitive to the environment. In other words, different genes underlie shape variation in different environments. These results have major implications for research aimed at identifying the genes that underlie species divergence. They suggest that results could be misleading if the experiments are not performed in the ‘correct’ environment, which is to say the environment in which evolutionary divergence occurred.
Natural selection acts upon phenotypic variation. This notion has not changed since the time of Darwin. Recent work in Dr Albertson’s lab highlights the importance of genes in generating variation, however it also underscores the immense significance of the environment. As the field moves forward, it will be necessary to incorporate both variables (genes and environment) into a common framework in order to gain a comprehensive and mechanistic understanding of how species evolve.
Meet the researcher

Dr Craig Albertson
Associate Professor Department of Biology
University of Massachusetts,
Amherst
USA
Dr Craig Albertson is an Associate Professor in the Department of Biology at the University of Massachusetts, Amherst, where he studies the development and evolution of complex morphologies, using the craniofacial skeleton in bony fishes as his main experimental model. Dr Albertson was awarded his PhD from the University of New Hampshire in 2002 for a thesis entitled Genetic Basis of Adaptive Morphological Radiation in East African Cichlid Fishes. He then went on to carry out postdoctoral research on zebrafish developmental genetics at the Forsyth Institute and Harvard School of Dental Medicine. He won the Ernst Mayr Award in Evolutionary Biology, is an NSF CAREER award grantee, has been the Keynote or Invited speaker at numerous conferences, and received the College of Natural Sciences Outstanding Teacher Award in 2016.
CONTACT
E: albertson@bio.umass.edu
T: (+1) 413 545 2902
W: https://www.bio.umass.edu/biology/about/directories/faculty/rcraig-albertson
LAB MEMBERS
Moira Concannon (OEB PhD student)
Dina Navon (OEB PhD student)
Douglas Calenda (MCB MS student)
LAB ALUMNI
Kara Powder (Postdoctoral Fellow 2011–2016, now Assistant Professor, Clemson University)
Yinan Hu (OEB PhD student 2009–2015, now Postdoctoral Fellow, URI) Kevin Parsons (Postdoctoral Fellow 2009–2012, now Lecturer, University of Glasgow)
Jim Cooper (Postdoctoral Fellow 2007–2011, now Assistant Professor, Washington State University, Tri Cities)
Nicole (Jacobes) McDaniels (PhD student 2007–2011, now Assistant Professor, SUNY Herkimer)
FUNDING
NSF
NIH
KEY REFERENCES
KJ Parsons, M Conith, D Navon, J Wang, I Ea, K Groveas, C Campbell, RC Albertson, Foraging environment determines the genetic architecture and evolutionary potential of trophic morphology in cichlid fishes, Mol Ecol., 2016, In press. DOI: 10.1111/mec.13801.
Y Hu, RC Albertson, Hedgehog signaling mediates adaptive variation in a dynamic functional system in the cichlid skull, Proc. Natl. Acad. Sci. USA, 2014, 111, 8530–8534.
KE Powder, H Cousin, G McLinden, RC Albertson, A non-synonymous mutation in the transcriptional regulator lbh is associated with cichlid craniofacial adaptation and neural crest cell development, Mol Biol Evol., 2014, 31, 3113–24.
KJ Parsons, AT Taylor, KE Powder, RC Albertson, Wnt signalling underlies the evolution of new phenotypes and craniofacial variability in Lake Malawi cichlids, Nat Commun., 2014, 5, 3629. KJ Parsons, RC Albertson, Unifying and generalizing evo-devo’s two strands, Trends Ecol Evol., 2013, 28, 584–591.


