Dr Gábor Balázsi – Rewiring DNA: Gene Circuits in Synthetic Biology
The futuristic field of synthetic biology has the potential to deliver exciting, innovative technologies – improved chemicals, materials, medicines, environmental solutions, and even smart biological devices – converging Mother Nature with human ingenuity. Dr Gábor Balázsi, of Stony Brook University, New York, is swapping silicon for DNA to create genetic circuits inside ‘programmable’ living cells. His research group is using synthetic gene circuits as research tools to study cell evolution, development, and cancer.
Not All Circuits Are Electrical
An organism’s phenotype is determined by proteins, the cell’s functional molecules. The production of proteins is determined by the genetic code – within the DNA genes. The genetic code consists of various combinations of four bases – A, T, C, and G – which encodes sequences of amino acids which join together to make proteins.
In gene expression, information flows from genes to RNA transcript molecules (a process known as transcription), and then from mRNA to protein production (translation). The ATCG genetic code is the biological equivalent to binary code in electronics – sequences of 1s and 0s. A major vision of synthetic biology is the ability to program the genetic code for a desired output, i.e., to create ‘biological computers’. Synthetic biology is a multidisciplinary field, borrowing principles from electronics, computer science, engineering, and traditional biology. Nowhere is this more apparent than with gene circuits.
In nature, organisms have developed systems for regulating gene expression. Regulating the level of gene expression – ‘turning the dial’ for high or low production of proteins – enables cells to cope with stress. Certain proteins can induce or repress the transcription of specific genes – thus some genes can increase or decrease the expression of others – giving rise to gene regulatory networks.
Synthetic biology allows us to build synthetic gene regulatory networks with novel functions that don’t exist in the natural world. Synthetic gene circuits can be described as artificial gene networks assembled by humans. Complex functions can be engineered with such gene circuits – inducible expression, oscillators, logic gates, and even memory devices! The output in some synthetic gene circuits can become the input (creating feedback) – thus the network controls itself perpetually. Such self-control can be designed as positive feedback (PF – self-activation) or negative feedback (NF – self-repression).
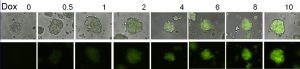
An example of tuning the levels of the GFP protein with Doxycycline (Dox; concentrations in ng/mL) in breast cancer cells using the Linearizer synthetic gene circuit. Credit Gábor Balázsi.
Dr Gábor Balázsi, Henry Laufer Associate Professor of Physical and Quantitative Biology at Stony Brook University, New York, is a pioneer in using network and systems approaches in biology. Gene networks and synthetic gene circuits are at the heart of his group’s research. A major part of this involves using synthetic gene circuits as perturbation tools, that is, applying precisely-controlled perturbations to cells in controlled environments and studying how the cells respond, or investigating how physical factors interact with known biological aspects of gene networks. Some of their most ambitious research has involved using synthetic gene circuits to create and evolve biological controller devices that mimic natural networks.
To have functionality, synthetic gene circuits must be inside a host organism – a ‘chassis’. The group’s first chassis of choice was that of yeast cells, known as Saccharomyces cerevisiae. The usefulness of yeast extends far beyond the making of bread and beer! It is a popular model organism for scientific research and biotechnology, and has several features that make it amenable to synthetic biology investigation. Yeast is easy to culture, easy to genetically manipulate, and its fully characterised genome resembles the human genome among others.
Gene circuits are assembled de novo. The individual genetic components are joined together to make a linear DNA construct and the construct is integrated into a circular DNA plasmid vector, allowing the transfer of genetic material into cells using standard molecular biology techniques. The artificial DNA is taken up by the yeast cells (or another host) in a process known as transfection. In this group’s studies, the DNA construct is integrated into the host cell’s chromosomal genome, whereas in studies by other groups, the synthetic circuit-containing plasmids can exist inside the cells independent of the chromosomes.
The cells are typically grown in liquid culture and subjected to various conditions. The output of the gene circuit is initially typically a fluorescent protein, to allow the researchers to test and troubleshoot circuit functionality. After culture, cells are normally subjected to flow cytometry and fluorescent microscopy. Flow cytometry is an ideal tool as it allows the circuit response in many individual cells to be measured quickly and automatically. It involves passing the cells single-file through a tube and measuring individual cell fluorescence.
‘We are looking at single cells and at genetic systems that we can dissect and understand gene by gene with a high level of detail,’ explains Dr Balázsi. The group has successfully transferred multiple synthetic gene circuits from yeast into mammalian cells – human cell lines. This is a major milestone, as it paves the way forward for medical applications of gene circuits.
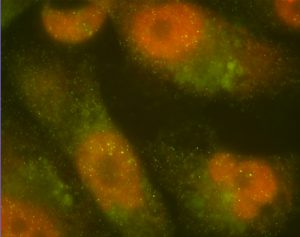
Metastatic breast cancer cells. Red and green mark a protein promoting and suppressing metastasis, respectively. Credit Gábor Balázsi.
Gene Networks in the Evolution of Cancer and Infection
Although we tend to think of evolution as occurring over millions of years, it can happen rapidly. Evolution plays a major role in rapidly augmenting the destructive cell populations that cause disease, including microbial infections and cancer. It is now recognised that evolution is not only driven by mutations in genes but also environment-dependent or random changes in protein levels. These changes happen largely by chance, stochastically or randomly, and genetically identical cells can have completely different phenotypes.
Disease-causing cells, including cancer cells and pathogens, start as a small group of genetically-identical cells, but then expand into a large phenotypically heterogeneous population. Some cells are stronger than others, and the strongest thrive. Surprisingly, cells that are strong in a given condition can be weak in another and vice versa. It is the various types of heterogeneity that are behind the disease’s resilience and tolerance to treatments – often, cancer tumours stop responding to chemotherapy, and likewise, infectious microbes can develop resistance to anti-microbials.
Dr Balázsi’s group has been investigating how cellular heterogeneity influences the evolution of cell populations, and the health implications of this. Dr Balázsi has recently been awarded a $1.8 million 5-year National Institutes of Health grant to search for the random triggers that turn benign cells into dangerous ones. ‘Cancer cells often don’t look the same in a matter of months, and drug-resistant microbes may not look the same in a matter of day,’ notes Dr Balázsi, who would like to know what causes these changes, and how we can prevent them.
Dr Balázsi hopes to make great strides in decoding how random genetic and protein-level changes occur and interact. A synergistic collaboration with Professor Marsha Rosner at the University of Chicago has already yielded ground-breaking insights. With cancer, cells divide rapidly to form a tumour. When a tumour grows large enough, it becomes metastatic and cancerous cells spread to other parts of the body. Metastatic tumours typically consist of a heterogeneous cell population but how this emerges is not fully understood. The findings from Drs Balázsi and Rosner demonstrate that the development of heterogeneous tumours and metastasis involve random changes in protein levels besides mutations. They hope that these insights will lead to more effective, safer cancer treatment regimes.
Developing Predictive Tools for Synthetic and Evolutionary Biology
A major theme in Dr Balázsi’s research is the power of prediction. In disease, as well as biotechnological applications, it is important to be able to predict a cell population’s evolutionary path – but evolution’s stochastic nature makes this difficult. Gene circuits can be inserted into industrial or future therapeutic bacteria, but will the cell-circuit system be functional in the long-term?
Supported by a 2009 NIH Director’s New Innovator Award, Dr Balázsi’s group has built computational models using fitness landscapes to predict the outcome of evolving cell populations. They then tested these predictions with ‘wet lab’ experiments involving yeast cell cultures containing a PF gene circuit under various toxin dose conditions. In some cultures, inducer molecules were added to activate the circuit. The circuit, when active, expresses a protein that mitigates the effect of the toxin, but the circuit’s activity is physiologically demanding on the yeast. When the toxin is present, the circuit’s activity is beneficial, but when absent, the circuit is potentially damaging to the cells.
The group found that when the costs outweigh the benefits, mutations within the module will deactivate it. When the benefits outweigh the costs, mutations within the circuit and within the yeast’s genome will jointly activate the module. Even in cultures lacking inducers, yeast still ‘learns’ to induce the circuit slightly through evolutionary selection in scenarios when this is beneficial – a remarkable finding! This work indicated that evolution can inadvertently switch genes and networks ‘on’ even when they are intended to be ‘off’ or vice versa – as can happen in disease. The group hopes to translate these findings into the development of more accurate predictive evolutionary models.

Fitness landscape and data from evolving the PF synthetic gene circuit. Evolution pushes cells towards the peaks on such landscapes. This means that expression has to drop in Dox (blue) and increase in Zeocin (red) for fitness to increase. Experimental measurements of gene expression (top timecourses) and fitness (bottom timecourses) confirm these expectations. Credit Gábor Balázsi.
Temperature Responses and DNA Dimming
Synthetic gene circuits could be used in ecological applications, but they would have to stand up to the elements. For instance, they may be inserted into crops to create tolerance against pathogens or drought – but will they work in spring and summer temperatures? Microbes containing gene circuits could clean toxic spills – but will they work under fluctuating temperatures? Temperature is a major factor in gene circuit performance, and Dr Balázsi’s group have investigated its influence.
In a recent study, they cultured yeast containing PF or NF circuits at cold, standard, and warm temperatures, and observed circuit responses with flow cytometry. They found that at non-optimal temperatures, cell-circuit systems make the choice between ‘switching on’ genes that confer stress resistance or stopping growth and protein synthesis. Moreover, temperature alterations influence the rates of biochemical reactions and protein structures. The findings suggest that at lower temperatures, achieving a certain gene expression level requires lower doses of inducer molecules – which could reduce the cost and environmental impact of reagents in industrial applications of synthetic biology.
Biotechnologists have long used ‘genetic switches’ to turn genes on or off, but a goal of synthetic biology is to intermediately adjust gene expression – much like a dimmer switch. In the late 2000s, in his previous post at University of Texas, Dr Balázsi led a group of researchers that achieved just that. They serendipitously engineered yeast with a ‘DNA dimmer’ and by adding varying doses of anhydrotetracycline (inducer) they found that they could precisely control production of green fluorescent protein. For example, doubling the anhydrotetracycline concentration made the cells glow twice as bright. Later they successfully transferred this dimmer gene circuit into human cancer cells. The ability to precisely control gene expression will be critical to future synthetic biology applications, such as therapies for genetic disorders.

Dr Gábor Balázsi and Dr Daniel Charlebois. Credit Gábor Balázsi
Looking to the Future
Dr Balázsi is a pioneer in synthetic gene circuits – manipulating biological systems similarly to how an electrical engineer may rearrange or optimise the components of an electronic circuit, with promising insights for future applications and disease. He now hopes to interface natural gene networks with synthetic gene circuits to study how controlling network dynamics affects cell phenotypes in cancer and microbial drug resistance.
Meet the researcher

Dr Gábor Balázsi
Henry Laufer Associate Professor
The Louis and Beatrice Laufer Center for Physical and Quantitative Biology
Associate Professor, Department of Biomedical Engineering
Stony Brook University
Stony Brook, NY
USA
Dr Gábor Balázsi is the Henry Laufer Associate Professor of Physical and Quantitative Biology at Stony Brook University, New York. He aims to develop a predictive, quantitative understanding of biological processes, including cellular decision making and the evolution of cell populations. Dr Balázsi is well-placed to lead his group with an impressive academic background in physics, biophysics, and systems biology. He graduated with BS Physics (1996) and MS Magnetism (1997), both from Babeş-Bolyai University of Cluj (Kolozsvár), the major center for Hungarian-language higher education in his native region of Transylvania, Romania. He then studied at the University of Missouri, St Louis, USA, where he gained his MS and PhD in Physics (1999 and 2001). In 2006, he took up an Assistant Professorship at the Systems Biology Department, University of Texas MD Anderson Cancer Center, before being promoted to Associate Professor in 2012. In 2014, he took up his current appointment.
CONTACT
E: gabor.balazsi@stonybrook.edu
W: https://www.stonybrook.edu/commcms/bme/people/g_balazsi.php
KEY COLLABORATORS
Professor Marsha Rosner (University of Chicago)
Professor Lanying Zeng (Texas A&M University)
FUNDING
2017: National Institutes of Health: Maximizing Investigators’ Research Award ($1.8 million, 5 years)
2009: National Institutes of Health: NIH Director’s New Innovator Award ($2.3 million, 5 years)
FURTHER READING
DA Charlebois, K Hauser, S Marshall, G Balázsi, Multiscale effects of heating and cooling on genes and gene networks, Proceedings of the National Academy of Sciences, 2018, 115, E10797–E10806.
J Lee, J Lee, KS Farquhar, et al, Network of mutually repressive metastasis regulators can promote cell heterogeneity and metastatic transitions, Proceedings of the National Academy of Sciences, 2014, 111, E364–73.
D Nevozhay, T Zal, G Balázsi, Transferring a synthetic gene circuit from yeast to mammalian cells, Nature Communications, 2013, 4, 1451.
D Nevozhay, RM Adams, E Van Itallie, et al, Mapping the environmental fitness landscape of a synthetic gene circuit. PLoS Computational Biology, 2012, 8, e1002480.
D Nevozhay, R Adams, K Murphy, et al, Negative autoregulation linearizes the dose response and suppresses the heterogeneity of gene expression, Proceedings of the National Academy of Sciences, 2009, 106, 5123–5128.

