Dr Jason Crawford – Could a Better Understanding of Bacteria Prevent Colorectal Cancer?
Our digestive system contains trillions of bacterial cells, constituting a highly diverse community of microorganisms living within us that can influence human physiology and cause disease. Dr Jason Crawford at Yale University has extensively researched some of the more harmful bacterial strains, looking at how they promote inflammation and colorectal cancer.
The Good, the Bad and the Ugly
Bacteria are literally everywhere. You only have to look as far as your hand to see some, albeit through a microscope rather than the naked eye. Whether they are on your dining table, on your person, or on your phone, avoiding bacteria is impossible.
That’s not to say that all bacteria are bad. Bacteria get a bad reputation – they may be one of the major reasons behind why we become ill, but they can also do a lot of good for our bodies. Almost 100 trillion bacterial cells are currently living in your gut. Now, there’s no reason to panic – most ‘good’ bacteria are vital to our continued survival. Not only do they help our bodies digest food and absorb nutrients, but they also produce vitamins in our intestinal tracts, including vitamin K needed for blood coagulation and vitamin B6 important for maintaining a healthy nervous system.
They can also help our immune systems out when responding to pathogens and harmful bacteria, by crowding them out in our gut, producing certain materials to prevent their growth and kickstarting our immune response into action.
 Getting to Know Your Belly
Getting to Know Your Belly
The types of bacteria that live in your gut can vary massively over time. From probiotics such as Lactobacillus, Bifidobacteria and Streptococcus, to other forms of microorganisms such as Bacteroides and Escherichia, your gut is home to what is known as ‘gut microflora’. Gut flora constitutes different bacterial species that vary in ratio and abundance according to time, diet and changes to overall health. The bacteria that live alongside us contain roughly 100 times more genes than the human genome itself representing a potentially huge number of biologically active small molecules that can be processed and synthesised by the bacteria living within their human hosts.
Small molecules produced by bacteria can affect both the numbers and types of microorganisms living within the gut, but they can also influence the physiology of their hosts by regulating a variety of different cellular processes.
Not much is known about how small molecules produced by bacteria influence human health and so their functions and how they are produced are an important area of biomedical research, especially as not all strains of bacteria in the gut are harmless. Dr Jason Crawford and his team at Yale University have been investigating the strains of gut bacteria that have the potential to cause disease and pose a harmful risk.
‘We at the Crawford Lab primarily focus on the molecular mechanisms of the interactions between humans and bacteria.’
Pondering Pathogen Potential
Dr Crawford and his team focus on the ‘chemistry at the human-bacteria interface’, predominantly looking at how bacteria regulate inflammation and cancer and how they could be exploited for human treatments. He describes how: ‘We primarily focus on the molecular mechanisms of the interactions between humans and bacteria. We specifically focus on how bacterial metabolism can regulate cancer initiation and treatment, immune regulation, and bacterial pathogenicity in the human host.’
The team has researched and developed a number of methods to ‘mine’ the genomic sequences of pathogenic bacterial strains, to determine how they cause disease. The genomic sequences of bacteria encode the structures of the small molecules that they produce in the body, which can lead to disease. The team can then look to utilise this knowledge by designing drugs capable of inhibiting disease processes.
Through their research, Dr Crawford and his team have characterised a number of unusual clusters of genes suspected of synthesising or manufacturing important small molecules. As he states: ‘These small molecules often regulate complex interactions with their animal hosts, hold a rich history of being utilised as human drugs, and serve as excellent molecular probes for identifying new drug targets for a wide variety of diseases.’ As such, researching and identifying the gene clusters responsible for producing small molecules of this kind is a major focus of Dr Crawford and his team’s work.

The Bad Side of E. coli
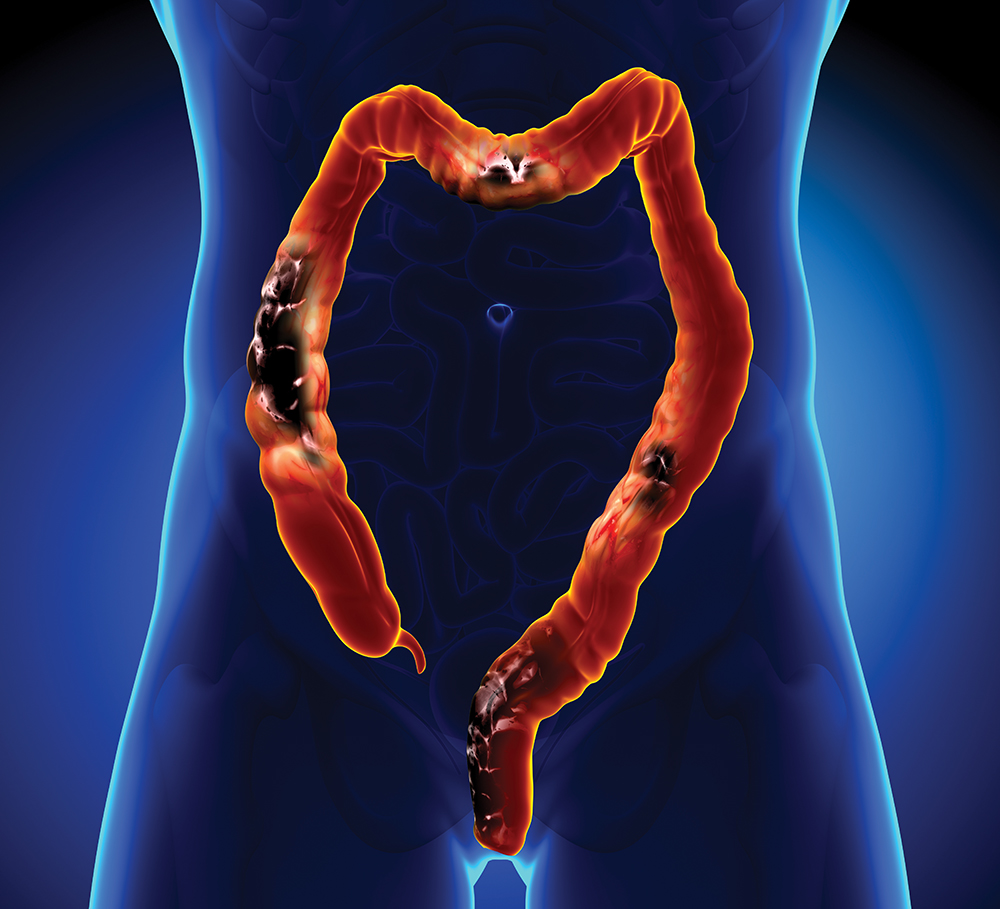
Over the past six years, Dr Crawford’s research team has focused a good portion of their research on gut bacteria called Escherichia coli (E. coli). E. coli is no stranger to the world of research, especially in babies and infants, as it is the first type of bacteria to colonise the gut after birth and is thought to live in about 90% of the human population.
It is usually a harmless type of bacteria. However, Dr Crawford’s team and his key collaborators study certain strains of E. coli that are linked to the onset of colorectal cancer – cancer of the intestines. Through their research, they have characterised a particular genetic pathway that encodes small biological molecules known as ‘colibactins.’ While its fully mature structure has been challenging to characterise, they have shown that fragments of the small molecule cause damage to DNA consistent with colibactin’s proposed role in causing colorectal cancer.
Introducing Colibactin
During their research, the team used a combination of techniques to characterise the structure of the ‘colibactin warhead’ – a critical component of the colibactin molecule responsible for damaging DNA and initiating tumour growth. The colibactin warhead participates in a process called DNA alkylation and cross-linking. DNA alkylation is a process where a specific section of a molecule (a reactive alkyl group) forms a bond with DNA.
DNA cross-links result from two bonds to the DNA duplex, locking DNA strands together. Modification of DNA with cross-links can be highly toxic, impairing cellular processes that use DNA as a template, such as the replication of DNA when cells divide or the production of RNA to produce proteins. Cross-links, for example, can induce multiple DNA repair mechanisms and can cause downstream DNA breaks in the double-stranded helix. The accumulation of DNA damage can lead to tumour formation.
The team’s research demonstrated that certain forms of model colibactins, closely related to natural colibactins, are more efficient at initiating DNA damage than others. The pathway that makes colibactin uses giant modular enzymes called nonribosomal peptide synthetases and polyketide synthases to synthesise precolibactins in an ‘assembly-line’ manner that are later converted into colibactins, through an enzyme called a peptidase, dedicated to the task. Dr Crawford and his team looked at the effects of inactivating individual modules of the assembly line through a novel characterisation process called domain-targeted metabolomics.
The process involves investigating the synthesis pathways bacteria use to produce small molecules by combining gene editing of the different parts of each enzyme with analysis of the pathway and the products produced. The team then investigated the different steps in the assembly line in more detail and identified two new precolibactin products that could also potentially cause damage to DNA. A similar technical approach can be used in the future to decipher the many steps in the synthesis pathways of important small molecules produced by bacteria with very high precision.
Gutsy Work
The team’s research has also looked at utilising the findings from their previous work to investigate potential molecules that could offer resistance against colibactin’s ability to cause colorectal cancer. To continue their work, the products created by the DNA gene cluster that produces colibactin need to be investigated further. As Dr Crawford states in one of his research papers: ‘Though advances toward elucidating (pre)colibactin synthesis have been made, the functions and mechanisms of several clb gene products remain poorly understood.’
Dr Crawford’s team has more recently been investigating the molecular function of ClbS – a gene product that confers resistance to colibactin toxicity. Because ClbS neutralises the colibactin warhead, it could potentially be developed to prevent colibactin from causing cancer in the future.
Gut bacteria provide a fascinating area of scientific research and a huge potential reservoir of undiscovered drug-type molecules. Through the research of Dr Crawford and his team, science is now one step closer to understanding, combatting and preventing the onset of cancer in the intestine, utilising the strains of bacteria that live in our gut to our advantage. As their work supports, maybe the treatments to some of life’s most debilitating conditions have been in the pit of our stomachs all this time.
Meet the researcher

Professor Jason M. Crawford
Yale University
West Haven, CT
USA
Professor Jason M. Crawford studied Biological Chemistry, graduating with a BS in Chemistry, with an emphasis in Biotechnology in 2001. He completed an MA in Bioorganic Chemistry in 2003, before receiving a PhD in Bioorganic Chemistry in 2008 at the Johns Hopkins University. He then went on to work as a Damon Runyon Cancer Research Foundation Postdoctoral Fellow at Harvard Medical School in the Department of Biological Chemistry & Molecular Pharmacology until 2012, before joining Yale University as an Assistant Professor of Chemistry and of Microbial Pathogenesis in 2012, where he has remained ever since. Jason is currently the Maxine F. Singer Associate Professor of Chemistry and of Microbial Pathogenesis, where he serves as a member of the Yale Chemical Biology Institute, the Comprehensive Cancer Center, and the Center for Pulmonary Infection Research & Treatment. He has also received a number of awards and honours along the way. In 2007, he received the Sarah and Adolph Roseman Achievement Award from the Johns Hopkins University. In 2011, he received the NIH Pathway to Independence Award at Harvard Medical School. At Yale University, he has received the Searle Scholars, the NIH New Innovator, the Dale. F Frey Breakthrough Scientist, the Damon Runyon-Rachleff Innovation, the Camille & Henry Dreyfus Teacher-Scholar, and the Burroughs Wellcome Investigators in the Pathogenesis of Infectious Disease (PATH) awards over the last six years.
CONTACT
E: jason.crawford@yale.edu
W: https://crawfordlab.yale.edu/
KEY COLLABORATORS
For the selected project, Jason Crawford is indebted to the following key collaborators.
Seth B. Herzon, Yale University, USA
Steven D. Bruner, University of Florida, USA
Christian Jobin, University of Florida, USA
FUNDING
NIH National Cancer Institute (1DP2-CA186575 & R01-CA215553)
Burroughs Wellcome Fund (1016720)
Camille & Henry Dreyfus Foundation (TC-17-011)
Damon Runyon Cancer Research Foundation (DRR-39-16)
Yale Comprehensive Cancer Center
FURTHER READING
A Healy, H Nikolayevskiy, J Patel, J Crawford and S Herzon, A Mechanistic Model for Colibactin-Induced Genotoxicity, Journal of the American Chemical Society, 2016, 138, 15563–15570.
E Trautman, A Healy, E Shine, S Herzon and J Crawford, Domain-Targeted Metabolomics Delineates the Heterocycle Assembly Steps of Colibactin Biosynthesis, Journal of the American Chemical Society, 2017, 139, 4195–4201.
M Vizcaino and J Crawford, The colibactin warhead crosslinks DNA, Nature Chemistry, 2015, 7, 411–415.
P Tripathi, E Shine, A Healy, CS Kim, S Herzon, S Bruner and J Crawford, ClbS Is a Cyclopropane Hydrolase That Confers Colibactin Resistance, Journal of the American Chemical Society, 2017, 139, 17719–17722.