Professor Jeffrey C. Hoch – A Box in the Clouds
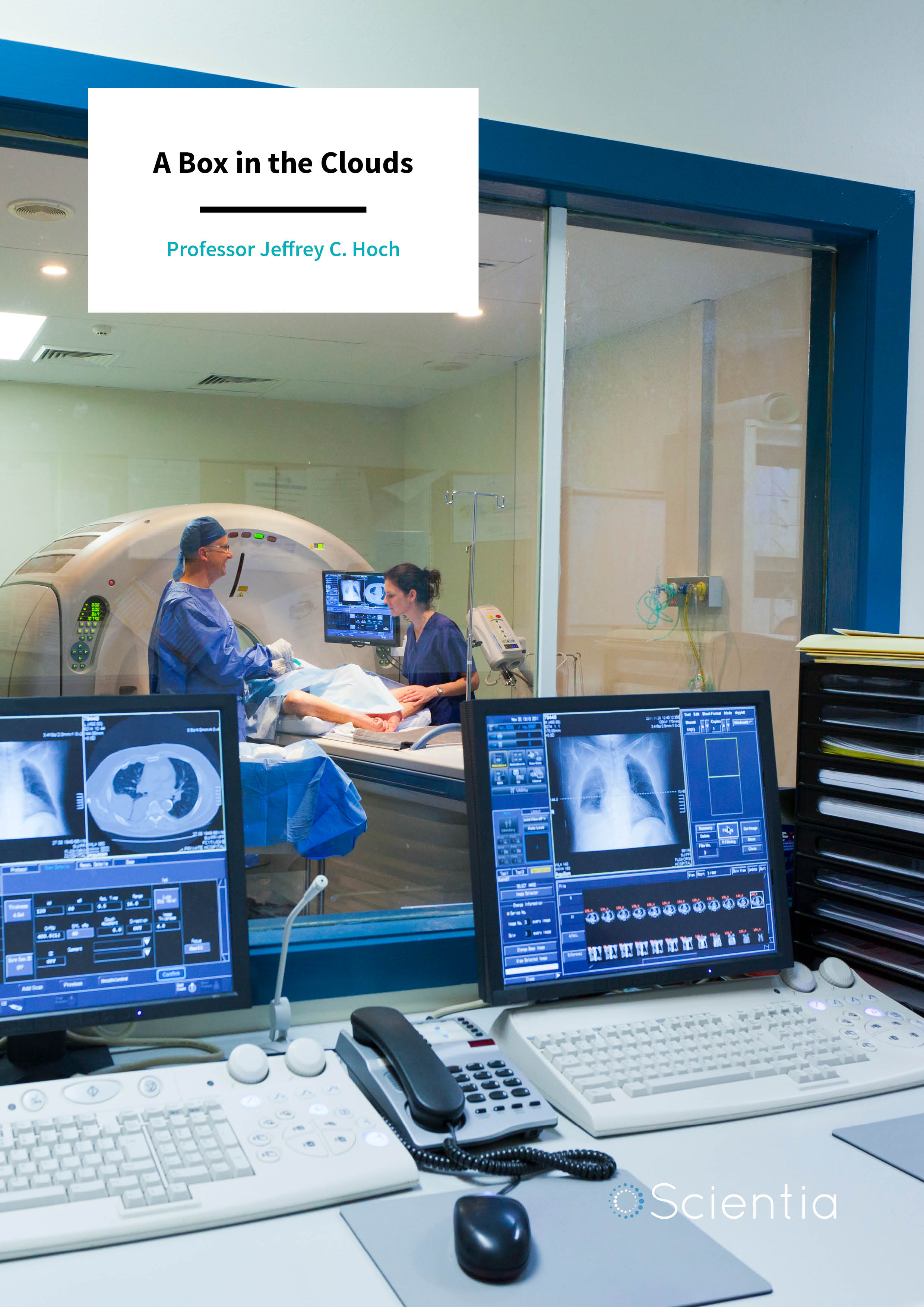
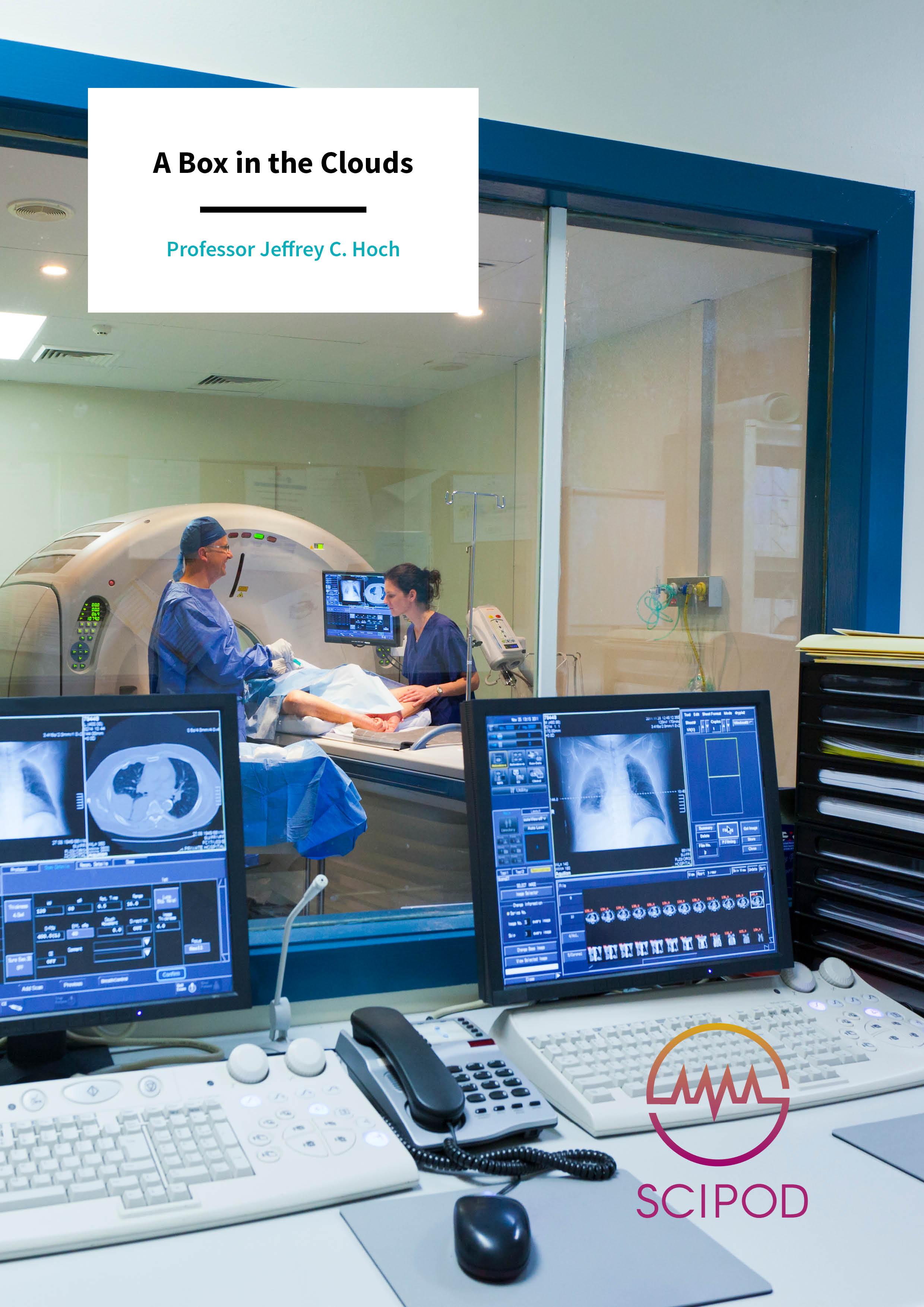
Nuclear magnetic resonance (NMR) is without doubt one of the most exciting analytic methods available in biomolecular medicine. Applications include structural biology, metabolic studies, disease diagnosis, and drug discovery. However, the use of NMR can be daunting and complicated, with a multitude of diverse computer programs for analysing the data to choose from. Professor Jeffrey C. Hoch from the University of Connecticut, USA, leads the development of the NMRbox platform, an extensive, freely available, not-for-profit resource aiming to help bring order to this chaos.
‘DNA is the blueprint of life’, goes the oft-repeated phrase. Yet few who hear it ever delve deeper to ask what that blueprint is actually for, to hear about proteins, the fascinating machinery, scaffolding, and sensors which are encoded by that string of information. Present in every cell, essential for life itself, with a dizzying range of structures, shapes and functions, proteins are the difference between the dead blueprint and the living factory.
Deconvoluted
This is, of course, an oversimplification. Atomic nuclei surrounded by a strong magnetic field will tend to take up an aligned magnetic direction, something which occurs in all atoms with odd numbers of protons and neutrons. Introducing a second, weaker magnetic field will disrupt that alignment, and doing so using an oscillating field will lead to transient pulses of disruption. The nuclei respond by emitting an electromagnetic signal, which can then be detected by the NMR machine. Most importantly, the frequency emitted is dependent on the atomic environment – a carbon nucleus bound to four hydrogens will emit a different signal to one bound to three hydrogens and a hydroxyl group. This is the attribute which allows NMR researchers determine chemical structures.
Proteins are complex molecules and thus the NMR signal from a protein will consist of a number of different signals mixed together. Pulling these signals apart to determine which belongs to which nuclei is a difficult process, requiring separation by mathematical techniques such as Fourier transforms (which convert a mixture of waveforms into discrete peaks) or maximum entropy reconstruction, which is often combined with sparse sampling (deliberately taking fewer readings than would normally be required). Nor is this enough to determine a structure. Thus, a researcher will need to use several different NMR methods (with delightful names such as COSY, NOESY and TOCSY) in order to fully determine a protein structure. This cross-checking and signal assignment will often take up the majority of a researcher’s time.
‘To set up a new NMR lab from scratch, you need months to find, assemble, and install the software, and technical expertise to maintain it.’
As may already be apparent, NMR is a complicated and heavily computer-dependent field of research, generating vast reams of data which require specialised software to interpret, visualise, and understand. This has led to a proliferation of software variants, most of which have been developed by academic groups in response to their own particular needs. Keeping an overview of this multitude of software options is made even more complex by development cycles, forking of projects to develop new variants, or the eventual discontinuation of development when the researchers graduate, retire, or simply run out of funding.
While this complexity is rarely a problem for those working on their own software, it does challenge those who follow after. Simply finding the ideal program is an ordeal in itself; scientific journals alone contain citations of hundreds of different software packages. To further complicate matters, the value of scientific discoveries increasingly rests on their ability to be reproduced, which in turn requires that the software used to make those discoveries is available for other researchers. Yet the range of software versions and oft-lacking archiving processes in academic laboratories means that software, particularly that used in older publications, is difficult to find.

Bringing Order to the Mix
This difficulty caught the attention of Professor Jeffrey C. Hoch of the University of Connecticut. Having been part of the NMR research field for over four decades, he had noticed the increasing difficulty which new entrants had with developing or identifying ideal software for their needs. ‘To set up a new NMR lab from scratch, you need months to find, assemble, and install the software, and technical expertise to maintain it’ comments Dr Hoch.

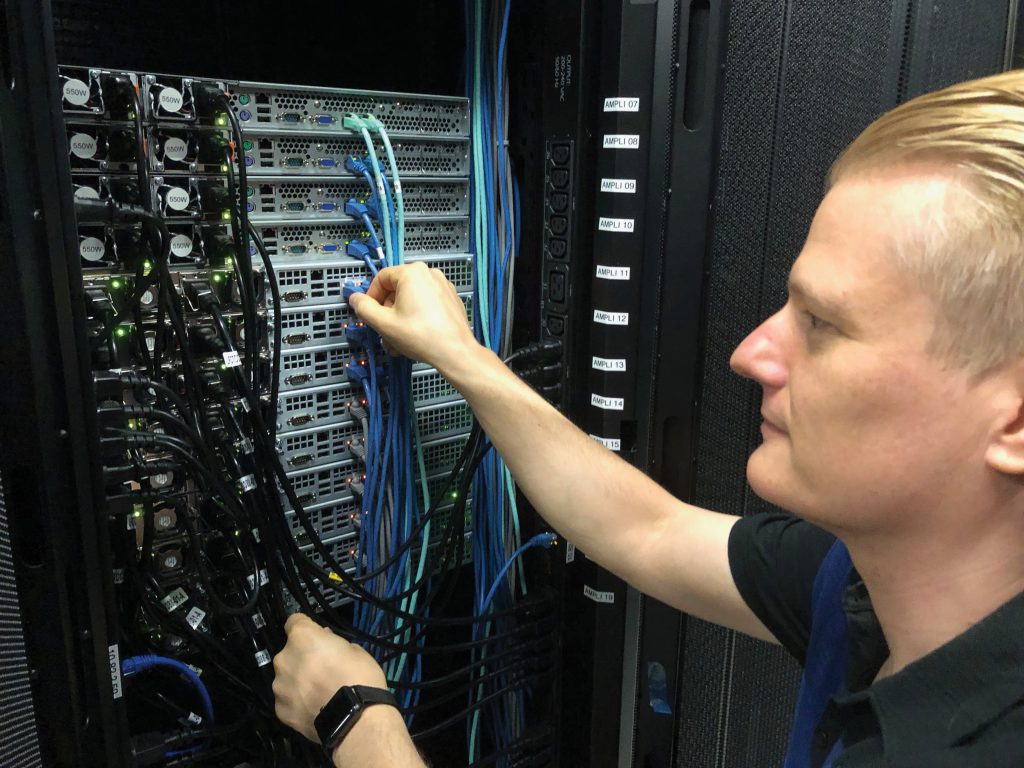
To help offset this steadily-increasing difficulty, Dr Hoch and his collaborators used a grant from the National Institutes of Health to establish the NMRbox, a set of virtual machines dedicated to the needs of NMR researchers. Hosted on a central server, it provides access to over one hundred NMR software packages which run the gamut from visualisation to validation. A detailed registry of all software currently available is provided, thus allowing newcomers to the NMR field to quickly find the ideal program. It also automatically links to protein structures and publications which have been generated using those programs to allow the comparison of different packages.
NMR work uses large data sets and requires computationally challenging interpretation to determine protein structures. This is often beyond the capabilities of academic laboratories, or requires long calculation times even if it is possible. NMRbox solves this problem by providing a ‘virtual machine’ for each researcher who logs in. This acts as a personal computer environment within the server in which they have access to all of the required software, yet the software itself runs on the high-performance equipment located in the server station. This essentially offloads the difficult work onto the dedicated (and expensive) equipment in the central location, while scientists simply need to download the results when they are available.
This centralisation of software also provides efficiency gains. The dedicated IT support frees scientists from the struggles of understanding why each program is refusing to work, while central upgrading processes ensure that the current version is always in play. This allows time-pressed researchers to focus directly on their work. Moreover, the NMRbox platform explicitly stores older versions of each package, allowing older papers to be accurately reproduced.
Server-based approaches are rapidly growing in popularity in all areas of computing – we simply consider the many things are now proudly advertised as being ‘in the cloud’. This has also proven true for NMRbox, which is going from strength to strength. The user-base now covers researchers from over 33 countries, with more expected in the coming year. This rapid rise in popularity has prompted the developers to double their computing capacity in the coming year, turning NMRbox into an essential part of any NMR research program.

The Magnet Gap
Although systems such as NMRbox allow researchers from across the world to examine their NMR data, it does not help with the act of gathering the data itself. For this, researchers need access to an NMR machine, a large and expensive piece of equipment which requires liquid nitrogen and helium to maintain powerful magnets and where the strength of the magnetic field puts a hard limit on the resolution which can be achieved. As Dr Hoch notes, ‘Sensitivity and resolution are key limiting factors in the application of NMR to challenging biomolecular systems, with higher magnetic fields improving both sensitivity and resolution.’
The next generation of NMR machines are already in development, working at 1.2 GHz, the equivalent of a magnetic field strength of 28 tesla (for comparison, the strength of a typical fridge magnet is around five milli-tesla). This exceptional magnetic strength means that the next generation will be able to detect far more interactions than ever before, particularly necessary for research into unstructured proteins or large complex biomolecules.
The world of research is a highly competitive one, with fierce rivalries not only between scientists but between nations. The level of funding and the equipment available to researchers within a particular region can spur the creation of a centre of excellence or force frustrated scientists to move abroad. These decisions connect the political and the scientific arenas, and are the source of much discussion. One of these factors is known as the ‘magnet gap’, the difference in regional availability for the upcoming generation of NMR machines. The instruments are extremely sensitive but also extremely expensive, and efforts to fund the next generation of machine within the United States have lagged those in other countries. At the same time, around ten instruments have been ordered by various locations within Europe.
Expert scientists within the USA are already beginning to worry about this. ‘In addition to the competitive disadvantage that investigators in the US will face when European scientists gains access to 1.2 GHz instruments,’ notes Dr Hoch, ‘students and trainees seeking access to state-of-the-art instrumentation will be forced to leave the US for training, resulting in a brain drain.’ This process has long been occurring from less-funded countries to research titans such as the USA and European Union – however the prospect of researchers fleeing the USA is an uncomfortable turn-around for scientists and politicians alike.

Implications for Machine Learning
Professor Hoch predicts that there is ‘an explosive growth of machine learning on the horizon.’ The study of how systems can automatically learn from data without being explicitly programmed to do so has huge and far-reaching implications for medical science. One potential barrier, however, is the lack of abundant training data, and here NMR lags behind other fields. There are public repositories of NMR data, one of which Professor Hoch is co-head (Biological Magnetic Resonance Data Bank – BMRB – based at the University of Wisconsin), but the amount of data deposited is minuscule compared to the amount of NMR data collected. As of yet, it is nowhere near the level of ‘big data’. One hope is that NMRbox will help make it easier for investigators to deposit their data in BMRB, thus taking forward this emergent field of research.
The Box of the Future
The NMRbox platform is already proving to be highly popular with researchers in the protein structure field. Yet Dr Hoch and his collaborators are not standing still, they intend to include yet more features into the system to assist with collaborative work and experimental reproducibility. With workflow metadata, processes for Bayesian inference and overarching systems to combine multiple software platforms, there is no lack of things to do.
Meanwhile Dr Hoch is wholly encouraging for all those who wish to get starting in NMR research via his platform system. Registration is free, he notes, so ‘users can kick the tires and figure out which method is best.’
Meet the researcher

Professor Jeffrey C. Hoch
Department of Molecular Biology and Biophysics
UConn Health
Farmington, CT
USA
Part of the NMR research scene for over four decades, Professor Jeffrey C. Hoch of the University of Connecticut is a well-established expert in this field. Starting with a PhD in physical chemistry from Harvard University, he has risen through the academic ranks to his current role in the Department for Molecular Biology and Biophysics. During this time, Professor Hoch has managed to appear on over 100 publications, pen several books, and accumulate a long list of invited talks and awards, while also fitting in the role of Director at the National Center for Biomolecular NMR Data Processing and Analysis. He is head of the NMRbox initiative.
CONTACT
T: +1 860-679-3566
KEY COLLABORATORS
The success of NMRbox is the result of close collaboration among teams based at UConn Health and the University of Wisconsin, and an active group of external collaborators.
UConn Health
Mark Maciejewski
Adam Schuyler
Michael Gryk
Ion Moraru
Yulia Pustovalova
Irina Bezsonova
Dmitry Korzhnev
Gerard Weatherby
University of Wisconsin
Pedro Romero
Hamid Eghbalnia
Eldon Ulrich
Miron Livny
External Collaborators
Frank Delaglio (US National Institute of Standards and University of Maryland)
Tatyana Polenova (University of Delaware)
David Rovnyak (Bucknell University)
Hari Arthanari (Harvard Medical School)
Robert Dempski (Worcester Polytechnic Institute)
Elizabeth Bafaro (Worcester Polytechnic Institute)
FUNDING
National Institutes of Health:
P41GM111135 (in support of NMRbox)
R01GM123249 (in support of signal processing research using NMRbox)
FURTHER READING
K Bourzac, How to Track Metabolites in Tissues Using NMR; The Scientist Magazine, 2018, 184.
MW Maciejewski, et al, NMRbox: A Resource for Biomolecular NMR Computation, Biophysical Journal, 2017, 112, 1529–1534.
JC Hoch, Beyond Fourier, Journal of Magnetic Resonance, 2017, 283, 117–123.
M Mobli, MW Maciejewski, AD Schuyler, AS Stern, JC Hoch, Sparse sampling methods in multidimensional NMR, Physical Chemistry Chemical Physics, 2012, 14, 10835–10843
M Mobli, JC Hoch, Nonuniform sampling and non-Fourier signal processing methods in multidimensional NMR, Progress in Nuclear Magnetic Resonance Spectroscopy, 2014, 83, 21–41.