Professor Naoko Tanese – Huntingtin: Its Role In Gene Expression
Professor Naoko Tanese and her research team at New York University School of Medicine investigate transcriptional and post-transcriptional gene regulatory pathways. Specifically, Professor Tanese is interested in identifying the post-transcriptional functions of the Huntington’s disease protein huntingtin and how they differ in the disease state.
Huntington’s Disease
Huntington’s disease is a rare, but devastating hereditary disease that causes the breakdown of nerve cells in the brain over time. The average age of onset is between 30 and 50 years; however, the disease can occur earlier or later. The result is a progressive decline in cognitive, psychiatric and motor abilities. Most literature reports motor impairments occurring early in the disease followed by cognitive/psychiatric dysfunction. Cognitive impairments include difficulty in prioritising and focusing, lack of impulse control and self-awareness, and difficulty learning new information. Memory can also be severely compromised. This decline is accompanied by the development of psychiatric symptoms, typically depression and anxiety, but other symptoms such as obsession, compulsion and irritability can also occur. Another hallmark of Huntington’s disease is the development of involuntary and unwanted movements in early stages of the disease. Clinical diagnosis occurs by the onset of motor symptoms. These movements start out as small, muscle twitches in the face and extremities that eventually progress to the rest of the muscles in the body, causing movements to become more rigid and difficult to initiate. Particularly aggressive forms of the disease display symptoms early in life. When Huntington’s disease develops before the age of 21, it is known as Juvenile Huntington’s disease or JHD. While very similar to the adult form, JHD differs from Huntington’s disease in that specific motor symptoms develop at different times during the progression of the disease and seizures can also occur. Some studies also suggest that JHD progresses more rapidly than the adult onset disease. While there are treatments available to manage the symptoms, there are currently no cures that prevent the decline associated with the progression of the disease.
Huntington’s disease is caused by an inherited mutation in the Huntingtin gene, HTT, that results in over 40 repeats of the amino acid glutamine in the encoded huntingtin protein (Htt). A typical huntingtin protein has between seven and 35 repeats. Huntington’s disease is an autosomal dominant genetic disorder, meaning that just one of the two genes that are inherited from one’s parents needs to be mutated for the disease to arise. HTT is widely expressed throughout the body and is essential for normal development. It has also been implicated in many cellular functions. In her latest project, Professor Naoko Tanese and her research team have discovered novel functions of Htt in gene expression. Understanding these new Htt functional roles and how they differ in the case of mutant Htt may reveal novel pathways and targets for future therapeutic interventions.
Post-transcriptional Regulation of Gene Expression
Gene expression is the process by which information encoded by a gene is used to synthesise a gene product, typically a protein. Protein synthesis occurs in three main steps: transcription, posttranscriptional processing, and translation. During transcription, an enzyme called RNA polymerase reads DNA of the gene and creates a copy called messenger ribonucleic acid (mRNA) or a transcript. The transcript then undergoes post-transcriptional processing to improve recognition of the mRNA by the translational proteins and remove any portions of the sequence that should not be translated. During translation, the transcript is decoded by a multi-protein structure called the ribosome to generate a string of amino acids known as a peptide. The peptide then folds into secondary and tertiary structures to form the protein.
Gene expression can be regulated by several factors at different steps in the protein synthesis process. Gene expression is regulated post-transcriptionally by RNA binding proteins or RBPs. Some RBPs interact with one end of the transcript called the 3’ untranslated region (UTR) to affect its stability or distribution in the cell in a number of ways. RBPs can influence gene expression by sequestering mRNA in processing bodies. Processing bodies (P-bodies) are small granules or aggregates of mRNA-degrading proteins in the cytoplasm. Through degradation or storing of mRNA, the P-bodies reduce the amount of protein that can be made and thus downregulate gene expression. Gene expression can also be regulated post-transcriptionally through mRNA localization, or transportation of the transcript to specific locations in the cell.
Localisation of mRNA is particularly important in nerve cells, or neurons. Neurons send signals to other neurons through a gap between the two cells called the synapse. This interaction is strengthened or weakened based on how much activity the synapse experiences over time. This change in strength is known as synaptic plasticity. Synaptic plasticity is vital for learning and forming new memories. Mounting evidence suggests that the transport and translation of mRNAs to the synapse contributes to synaptic plasticity by rapidly replenishing proteins to branches of the neuron that are far away from the cell body, especially following synaptic transmission.
Htt in RNA Transport and Translation
Professor Tanese and her research team have extensive experience in transcriptional and post-transcriptional gene regulation. More recently, they have turned their attention to the function of Htt. Because the normal function of Htt and the mechanism by which its mutant counterpart contributes to Huntington’s disease is still largely unknown, they became increasingly interested in the role of Htt in post-transcriptional gene regulation as a novel approach to understanding the function of Htt normally and in the disease state.
The team began their investigation by determining where the Htt protein was located in neurons. Using advanced imaging and microscopy techniques, the research team discovered that Htt could be found near neuronal RNA granules and P-bodies. RNA granules are large RNA-protein complexes responsible for transporting mRNA to specific areas in the cell. To determine whether Htt influences mRNA localisation, the research team reduced the level of Htt in neurons grown in culture and examined its effect on transcript transportation. They found that the reduction of Htt in the cells disrupted mRNA localisation, suggesting that Htt may contribute to the structure of RNA granules during RNA transport.
To obtain a better picture of the processes that Htt is involved in and how they may differ for mutant Htt, Professor Tanese and her team identified proteins that interact with each Htt form. By identifying the primary functions of the proteins that interact with Htt, the team could gain a better understanding of the processes that Htt may influence. Analysis of Htt protein binding partners revealed that both normal and mutant Htt interact with proteins related to RNA metabolism and protein synthesis, including complexes involved in translation. ‘It is difficult to tease out various functions of Htt in this context,’ Professor Tanese explains, ‘but we have evidence supporting its role in RNA metabolism.’ The mutant Htt associated with RBPs and translation factors more than the normal Htt, suggesting that dysregulation of post-transcriptional gene regulation and translation may contribute to the development of the diseased state in Huntington’s disease.
Now that the research team knew that Htt interacted with translational proteins, they wanted to determine the function of the association between Htt and these proteins. To determine its function, the team examined translation in cells with normal and high concentrations of Htt. They discovered that translation was inhibited in cells with a higher concentration of Htt. Taken together with additional experiments the data support a role for Htt in mRNA transport and translation.
‘Understanding the mechanism of disease is critical to identifying targets for therapeutic intervention and treatment especially for HD for which there is no cure’
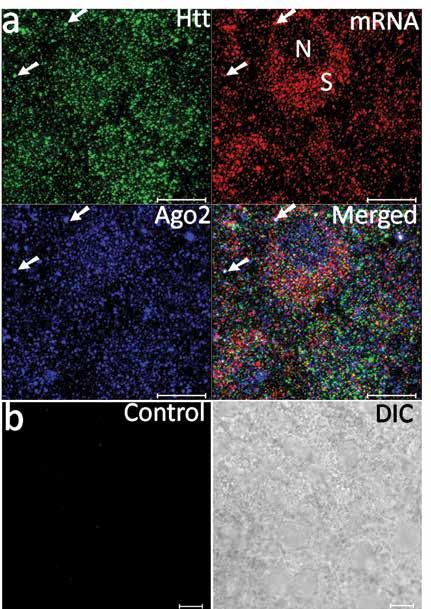
Visualisation of BDNF mRNA with huntingtin and argonaute proteins in the cortex of rat brain. Ma B, Culver BP, Baj G, Tongiorgi E, Chao MV and Tanese N, Mol Neurodegener., 2010, 27, 22.
Htt Self-Regulation
During their investigation, Professor Tanese and her team noticed that, in addition to proteins, Htt also associates with mRNA for ß-Actin – a protein essential for cell structure and rearrangement. This led them to ask whether Htt post-transcriptionally regulates gene expression through interaction with mRNAs and if so, what mRNAs did Htt regulate. To answer these questions, the research team strove to identify all other mRNAs that the protein associates with using a technique called RNA immunoprecipitation followed by RNA-Sequencing. To their surprise, they found that both the normal and mutant Htt protein strongly associated with their own HTT mRNA. The function of this association was investigated further by determining whether Htt regulates its own expression by inhibiting its translation through interactions with the HTT 3’ UTR. This is not uncommon, as many other proteins, such as TDP-43 (a protein whose mutation leads to the development of ALS, another neurodegenerative disease), regulate their own mRNA through interactions with the 3’UTR sequence. Professor Tanese found that when she linked another gene to the HTT 3’ UTR, there was a decrease in the linked protein’s expression level. Increasing the level of Htt protein further decreased the expression of the linked protein while reducing the Htt concentration had the opposite effect and increased the linked protein’s synthesis. This series of experiments provided strong evidence that Htt does in fact regulate translation of the HTT mRNA via the 3’ UTR sequence.
‘I am passionate about basic science research in which we strive to define molecular pathways and players that sustain cell homeostasis’
Mutant Htt and Mis-spliced mRNA
Examination of the huntingtin protein’s association with its own mRNA also revealed that mutant Htt associates with a mis-spliced HTT mRNA. Splicing is a common post-transcriptional modification that mRNA transcripts undergo to remove unnecessary strings of sequence. During the process, RBPs recruit proteins to remove the extraneous sections of RNA (introns) and splice together the remaining sections (exons). The RNA can be spliced in several different ways to produce multiple transcripts from one gene. The mis-spliced HTT mRNA has been incorrectly spliced resulting in a truncated protein encoded by exon 1. The fact that this association only occurs in the presence of mutant Htt and not normal Htt brings up the possibility that it may play a role in development of Huntington’s disease. Understanding the function of this truncated mRNA and its relationship to the mutant Htt could reveal a new target pathway for future Huntington’s disease treatments.
 Dissecting Htt’s Post-transcriptional Functions
Dissecting Htt’s Post-transcriptional Functions
Professor Tanese continues to tease out the role of Htt in the regulation, transport and translation of its own normal and mis-spliced mRNA. Currently, her lab is investigating the relationship between the mutant Htt and the mis-spliced mRNA further by identifying the mRNA sequence that allows them to associate, identifying any other proteins that may be involved in the interaction, and determining whether the 3’ UTR sequence of the mis-spliced mRNA has a function similar to its function in the normal HTT mRNA by regulating its translation.
In addition to its own mRNA, the research team confirmed that Htt associates with other mRNA. This led them to hypothesise that Htt may regulate the transport and translation of mRNA that are vital to neuron survival. Upcoming studies by this group will focus on identifying the other mRNAs that associate with Htt and mutant Htt during translation, the function of Htt in these relationships and how it differs between the normal and mutant protein. ‘In addition to investigating the nature of Htt protein – Htt mRNA interaction, we would like to identify other mRNAs selectively targeted by normal and mutant Htt protein.’ Professor Tanese explains. ‘We want to test the hypothesis that they have a role in the survival of neurons in the striatum, a brain region most susceptible to mutant Htt toxicity. We shall see what new direction the next discovery will take us.’
The team has uncovered novel roles for the normal and mutant Huntington’s disease protein huntingtin in post-transcriptional gene regulation and translation in neurons. These novel functions have several implications for the development of Huntington’s disease. Through understanding how Htt supports neurons with these novel functions, she and her team could reveal more accurate and effective pathways to target for Huntington’s disease treatment development.
Meet the researcher

Professor Naoko Tanese
Professor of Microbiology
Associate Dean for Biomedical Sciences
Director, Sackler Institute of Graduate Biomedical Sciences
New York University School of Medicine
New York, USA
Professor Naoko Tanese is a Professor of Microbiology, Associate Dean for Biomedical Sciences and Director of the Sackler Institute of Graduate Biomedical Sciences at New York University School of Medicine. She received her undergraduate degree in Chemistry at the University of Chicago in 1981. She went on to obtain her masters and PhD in biochemistry at Columbia University. She continued her postdoctoral training in biochemistry at the University of California, Berkley. She then made the transition to quite a different field of research, and now focuses on transcriptional and post-transcriptional gene regulatory pathways with an emphasis on the role of the huntingtin protein in these pathways and its influence on Huntington’s disease pathogenesis. In addition to her research, Professor Tanese is highly dedicated to student mentorship. In 2014, she was appointed Dean of the Sackler Institute of Graduate Biomedical Sciences. Her responsibilities include recruiting prospective students to the graduate program and providing oversight in the PhD training process.
CONTACT
T: (+1) 212 263 8945
W: www.med.nyu.edu/microbiology-parasitology/faculty/tanese-labmicrobiology
FUNDERS
National Institutes of Health Grant R01 NS061917
CHDI Foundation, Inc.
REFERENCES
Dunah AW, Jeong H, Griffin A, Kim Y-M, Standaert DG, Hersch SM, Mouradian MM, Young AB, Tanese N, Krainc D, Sp1 and TAFII130 transcriptional activity disrupted in early Huntington’s disease, Science, 2002, 296, 2238–43.
Savas, JN, Makusky A, Ottosen S, Baillat D, Then F, Krainc D, Shiekhattar R, Markey SP, Tanese N, Huntington’s disease protein contributes to RNA-mediated gene silencing through association with Argonaute and P-bodies, Proc. Natl. Acad. Sci. USA, 2008, 105, 10820–5.
Savas JN, Ma B, Deinhardt K, Culver BP, Restituito S, Wu L, Belasco JG, Chao MV, Tanese N, A role for Huntington’s disease protein in dendritic RNA granules, J. Biol. Chem., 2010, 285, 13142–53.
Culver BP, Savas JN, Park SK, Choi JH, Zheng S, Zeitlin SO, Yates III JR, Tanese N, Proteomic analysis of wild-type and mutant huntingtin associated proteins in mouse brains identifies unique interactions and involvement in protein synthesis, J. Biol. Chem., 2012, 287, 21599.
Culver BP, DeClercq J, Dolgalev I, Yu M-S, Ma B, Heguy A, Tanese N, Huntington disease protein huntingtin co-purifies with its own mRNA in mouse brain, J Huntingtons Dis., 2016, 5, 39 –51.