Professor Tony Moore – An Enzyme for the Future
Professor Tony Moore and his team at the University of Sussex are investigating the alternative oxidase. This enzyme provides an alternative route for molecular oxygen in the pathways of energy production that are often considered to be wasteful of energy but hold the key to the development of many different novel therapeutic treatments.
Energy is vital to every aspect of life. We as humans require it to perform everyday bodily functions, from breathing to keeping our hearts pumping, relying on incredibly complex mechanisms to furnish our energy demands. One of these mechanisms, called the respiratory chain is utilised by all eukaryotic cells in multicellular organisms such as plants and animals to produce energy.
In order to use this mechanism effectively, eukaryotic cells such as those within our own bodies contain mitochondria – tiny double-membranebound compartments called organelles that function as the powerhouse of the cell. Within the mitochondria is a convoluted and complicated folded inner membrane that houses the electron transport chain – this is the site of respiration and energy production. It is through this electron transport chain that mitochondria harness energy in the form of a simple molecule known as adenosine triphosphate (ATP).
 However, the mechanism through which this occurs is highly complex and relies on several proteins within the mitochondrion’s inner membrane to function correctly. In essence, the production of energy in all eukaryotic cells relies on a series of oxidationreduction reactions that eventually result in the addition of hydrogen to oxygen, thereby reducing it to water.
However, the mechanism through which this occurs is highly complex and relies on several proteins within the mitochondrion’s inner membrane to function correctly. In essence, the production of energy in all eukaryotic cells relies on a series of oxidationreduction reactions that eventually result in the addition of hydrogen to oxygen, thereby reducing it to water.
In doing so, the energy released during these oxidation-reduction reactions is used to ‘pump’ protons, positively charged hydrogen ions separated from their associated electrons, across the membrane thereby generating a proton electrochemical gradient – with a high concentration of protons on the outer side of the membrane and a low concentration on the inner side of the mitochondrial membrane.
Like water flowing downstream the protons are driven to follow the gradient and as the protons move back from the outer side (with a high concentration of protons) to the inner side (with a low concentration of protons), the energy released during this process is used to drive a molecular motor called the ATP synthase, resulting in the release of ATP for cellular activities. Multienzyme complexes located in the inner mitochondrial membrane mediate this reaction to release the energy that all complex cellular organisms then utilise to power their cellular activities and hence survive.
‘A key challenge in understanding how to control the alternative oxidases in plants, fungi and parasites rests upon the identification of the substrate and inhibitor-binding site(s) in this protein.’
Respiration
The process of respiration or energy production through the electron transport chain starts when the multi-enzyme complexes, NADH and succinate dehydrogenases (called complexes I and II respectively) transfer electrons into a pool of ubiquinone molecules and on to a multi-enzyme complex (the cytochrome bc1 complex) that then oxidises the pool and transfers the electrons to the final enzyme, known as cytochrome c oxidase. It is this protein which then reduces oxygen into water.
During each of the steps in the process, as electrons flow ‘downhill’ along the chain, free energy is released. It is this free energy that is used to form the proton gradient which drives the production of ATP via the ATP synthase – this is the source of energy that can be used to fuel vital biological processes required for cellular activity.
 Looking at the Alternatives
Looking at the Alternatives
While this electron pathway may be the most well-established route for the production of energy, further research by Professor Moore’s laboratory has found that alternative respiratory pathways also exist in plants, some fungi and parasites. This is due to the expression and presence of a single protein, known as the alternative oxidase (AOX). This enzyme, which is also located in mitochondria, enables plants, some fungi and parasites to bypass the cytochrome bc1 complex and cytochrome oxidase activities thereby releasing the energy as heat rather than forming a proton gradient.
This is a particular concern in terms of fungicide resistance, as many commercial fungicides target the respiratory chain at the level of the cytochrome bc1 complex thereby blocking the fungi’s ability to respire. If the alternative oxidase is present, however, it by-passes the site of fungicide action and hence enables the fungus to continue to respire and grow. Understanding the structure of AOX and how it works is, therefore, an important area of scientific research as it may provide information on an alternative target for the development of novel fungicides.
Professor Anthony Moore at the University of Sussex has dedicated over 40 years of his career to investigating the structure and function of the AOX enzyme in both plants and parasites. He describes how, ‘the Moore Lab’ is particularly interested in how the structure of this important but enigmatic protein influences its function in plants, fungi and parasites. Although this protein has been known about for over 100 years, relatively little is known about its mechanism of action or physiological significance.’
One of the consequences of alternative oxidase activity as indicated above is to facilitate the dissipation of energy as heat which some plants use to volatilise ‘smelly compounds’ to support pollination or to enable the plant to ‘burst through snow’ in order to flower during the winter months.
The AOX pathway can also be called into action in response to stresses such as drought, cold, harmful reactive oxygen species (ROS) and infection. Understanding the mechanism by which the protein is expressed and how it responds to these stresses have been particularly important to Professor Moore and his team, who have also recently identified AOX in human fungal pathogens, in addition to plants.

The Structure of Drug Development
Professor Moore and his team, in collaboration with Professor Kiyoshi Kita at the University of Tokyo and Nagasaki, have also been investigating the enzymatic structure of AOX within the protozoan parasite Trypanosoma brucei, known to cause African sleeping sickness.
The reason this parasite has been of such interest to Professor Moore and his collaborators in Japan is because of its total dependence, in the bloodstream form, on the alternative oxidase pathway to respire. Because this protein is absent from humans, understanding the nature of this alternative oxidase offers an attractive target for the design of new medicines to kill the parasite by disrupting its ability to survive.
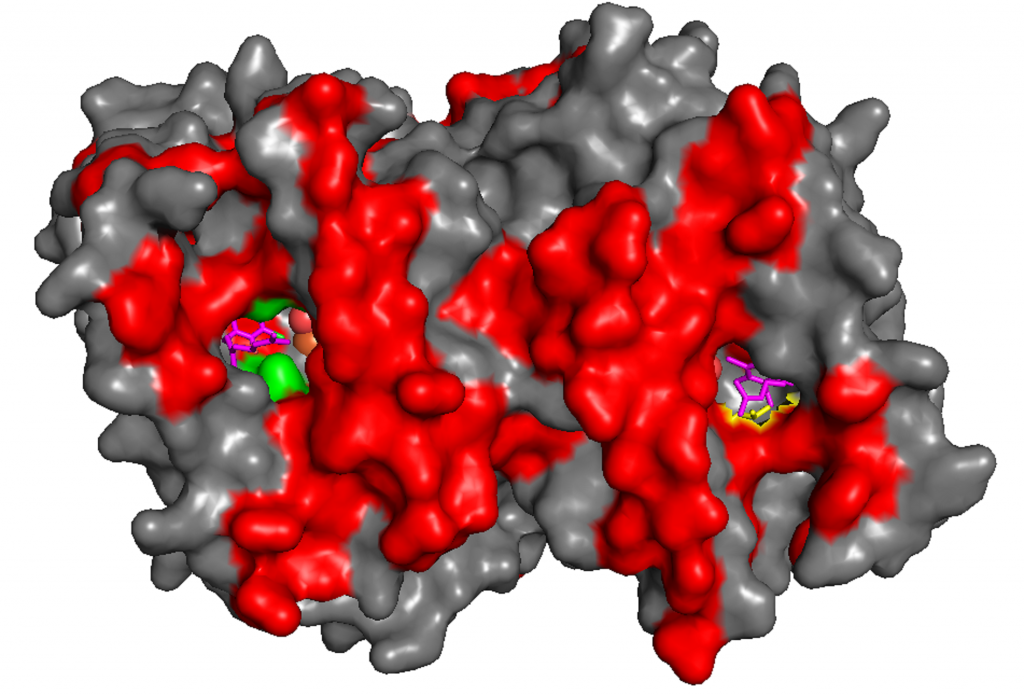
Recently, the Japanese and UK team determined the structure of the AOX at atomic level resolution both in the presence and absence of a number of specific AOX inhibitors. This has enabled them to identify the substrate and inhibitor binding site of the trypanosomal AOX. Such information is proving critical in the synthesis of novel inhibitors and effective drugs to hopefully treat trypanosomiasis or sleeping sickness.
Professor Moore states that, ‘a key challenge in understanding how to control the alternative oxidases in plants, fungi and parasites rests upon the identification of the substrate and inhibitor-binding site(s) in this protein.’ The substrate-binding site is the site at which the enzyme binds ubiquinone and possibly inhibitor molecules and so knowing more about the structure of this site and how AOX might bind to inhibitors will enable targeted drugs to be produced that can control its action.
Current and Future Research
In other research, Professor Moore and his team, have analysed the structures of AOXs from different species and in a review released last year, explained how differences in the response and sensitivity to specific AOX inhibitors could be related to structural differences in the channel within the protein which leads to the substrate binding site. They suggest that such structural differences indicate that all AOX proteins are not the same and hence such knowledge is critical for future structurebased drug design.
One particular area that such information has proven critical has been in the development of novel drugs to treat candidiasis. In addition to parasites such as trypanosomes, individuals with a compromised immune system are particularly susceptible to opportunistic diseases such as cryptosporidiosis, candidiasis and those caused by the microsporidia parasites. Examples of human fungal organisms that produce candidiasis include Candida albicans and Candida auris.
Candida auris is an emerging multi-drug resistant human fungal pathogen, which is resulting in a world-wide candidaemia epidemic of global concern. Its genome has been sequenced and Professor Moore’s group have confirmed that it does indeed contain a gene encoding the alternative oxidase. Independent testing of their current family of inhibitors has provided some spectacular results since some of Professor Moore’s inhibitors are proving effective at specifically inhibiting the growth and development of Candida auris. This is a major breakthrough as Candida auris is currently resistant to the usual drugs used to treat candidiasis.
Professor Moore is also collaborating with Dr Marten Szibor and Professor Howy Jacobs in Finland on the possible role the AOX protein could have in alleviating sepsis – a condition caused by an inflammatory response to infection that can lead to multiorgan dysfunction. The condition causes over 44,000 deaths per year, but his collaborator’s early results show a lot of promise.
Through the administration of AOX protein, a major symptom of sepsis known as septic shock, could possibly be protected against. This is thought to be due to the acceleration of the recovery process as a direct result of stimulated mitochondrial activity due to the presence of AOX. The promising and innovative nature of this work also shows potential in the effective design and development of novel AOX proteins to treat Alzheimer’s and Parkinson’s disease where changes in mitochondrial activity have been associated with the disease.
Professor Moore’s team has proved the importance of investigating these fundamental metabolic mechanisms. AOXs are now showing substantial promise in providing therapeutic support for a number of health conditions – an outcome spearheaded by Professor Moore’s ongoing dedication to understanding this enigmatic enzyme.
Meet the researcher

Professor Tony Moore FRSB, FLS
Biochemistry & Biomedicine
School of Life Sciences
University of Sussex
Brighton
UK
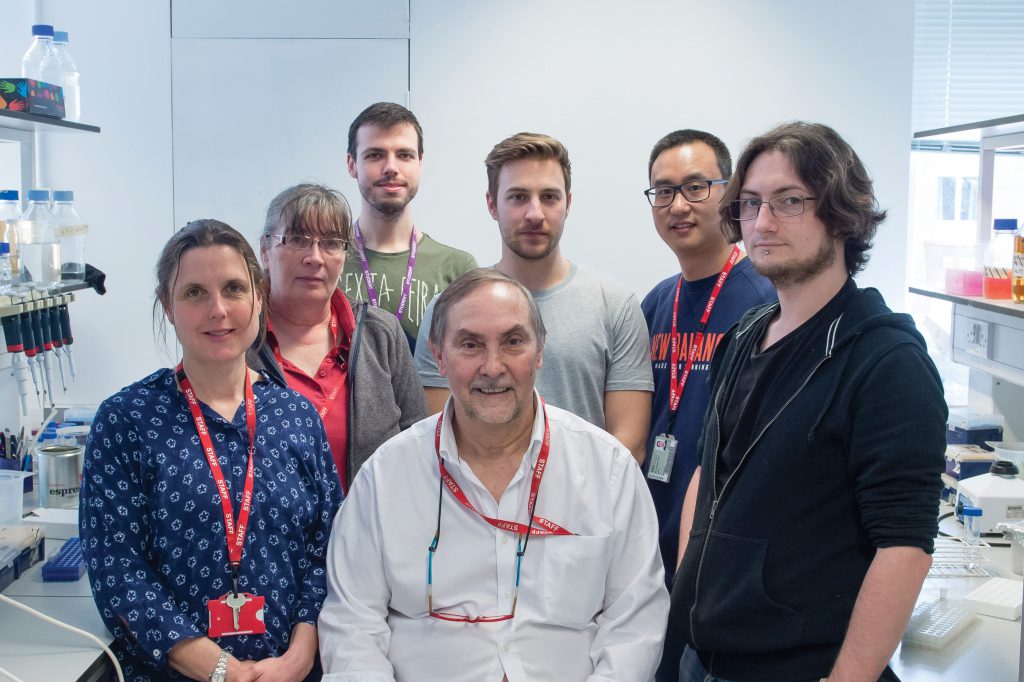 Professor Anthony (Tony) Moore is Professor of Biochemistry at the University of Sussex. He first developed an interest in mitochondria whilst at Shell Research Ltd, prior to undertaking a PhD in the same area at the University of Aberdeen. He later went on to study mitochondria at institutes such as the Johnson Research Foundation at the University of Pennsylvania, before joining the University of Sussex in 1979. Throughout his career, he has made many significant landmark discoveries into the understanding of alternative pathways of electron transport. He and his team at the Moore Lab are currently looking into the function, mechanisms and structure of alternative oxidases (AOX), enzymes which provide an alternative respiration pathway to plants, fungi, parasites and prokaryotes. They are working in collaboration with industrial and clinical partners to develop novel fungicides and therapeutics based on their research. In 2018, Professor Moore was selected as a finalist for the BBSRC Innovator of the Year Award and is currently in the process of spinning out a company called ‘AOX Technologies Ltd’ to commercialise his fungicides.
Professor Anthony (Tony) Moore is Professor of Biochemistry at the University of Sussex. He first developed an interest in mitochondria whilst at Shell Research Ltd, prior to undertaking a PhD in the same area at the University of Aberdeen. He later went on to study mitochondria at institutes such as the Johnson Research Foundation at the University of Pennsylvania, before joining the University of Sussex in 1979. Throughout his career, he has made many significant landmark discoveries into the understanding of alternative pathways of electron transport. He and his team at the Moore Lab are currently looking into the function, mechanisms and structure of alternative oxidases (AOX), enzymes which provide an alternative respiration pathway to plants, fungi, parasites and prokaryotes. They are working in collaboration with industrial and clinical partners to develop novel fungicides and therapeutics based on their research. In 2018, Professor Moore was selected as a finalist for the BBSRC Innovator of the Year Award and is currently in the process of spinning out a company called ‘AOX Technologies Ltd’ to commercialise his fungicides.
CONTACT
E: A.L.Moore@sussex.ac.uk
W: http://www.sussex.ac.uk/lifesci/moorelab/
KEY COLLABORATORS
Professor Kiyoshi Kita, Nagasaki University
Dr Marten Szibor, University of Tampere
Professor Howy Jacobs, University of Tampere
Dr John Misselbrook, MD Agform Ltd, UK
FUNDING
Biotechnology and Biological Sciences Research Council (BBSRC)
Wellcome Trust
FURTHER READING
B May, L Young and A Moore, Structural insights into the alternative oxidases: are all oxidases made equal? Biochemical Society Transactions, 2017, 45, 731–740.
A Moore, T Shiba, L Young, S Harada, K Kita and K Ito, Unraveling the heater: new insights into the structure of the alternative oxidase, The Annual Review of Plant Biology, 2013, 64, 637–663.



