Dr Kristen Kroll – New Models to Understand the Intricacies of Neurodevelopmental Disorders
Neurodevelopmental disorders (NDDs) are a complex group of diseases that profoundly impact the human population, emerging during brain development but often affecting individuals throughout their lives. Human models of NDDs are needed, as many aspects of both the human genome sequence and brain development are human-specific and not recapitulated in animal models. Dr Kristen Kroll, in the Department of Developmental Biology at Washington University School of Medicine, has spent her career modelling neural development and identifying how its disruption can contribute to NDDs.
Neurodevelopmental Disorders
Autism spectrum disorder (ASD), attention-deficit/hyperactivity disorder, and intellectual disability are examples of neurodevelopmental disorders (NDDs). According to the Centers for Disease Control and Prevention, approximately 1 in 54 children in the USA were identified as having ASD in 2016. Critically, data suggest that the prevalence of NDDs is increasing globally, while NDDs are diagnosed in individuals encompassing all countries, ethnicities, and socioeconomic backgrounds, and thus have an increasingly concerning impact on global health (World Health Organization).
Despite the widespread prevalence of NDDs, the causation of these disorders is complex and poorly understood, often involving disruption of multiple aspects of brain development. Thus, the main focus of Dr Kristen Kroll’s research at Washington University School of Medicine is studying embryonic development, with a particular focus on understanding how development of the human brain is regulated and on determining how dysregulation of this process gives rise to NDDs.
In the human brain, the cerebral cortex is a layered structure comprised of neurons responsible for mental functions including attention, awareness, perception, thought, memory, language, and reasoning. This structure begins forming during embryonic development through a complex process that must be precisely regulated. Brain development begins as some cells of the early embryo are directed to form the neural plate, with one end of this structure developing into the forebrain, from which the cortex is derived. Research in Dr Kroll’s laboratory focuses on understanding how this process is controlled and identifying the regulatory networks involved. Understanding these aspects of normal brain development is essential for determining how they are disrupted to cause human NDDs.
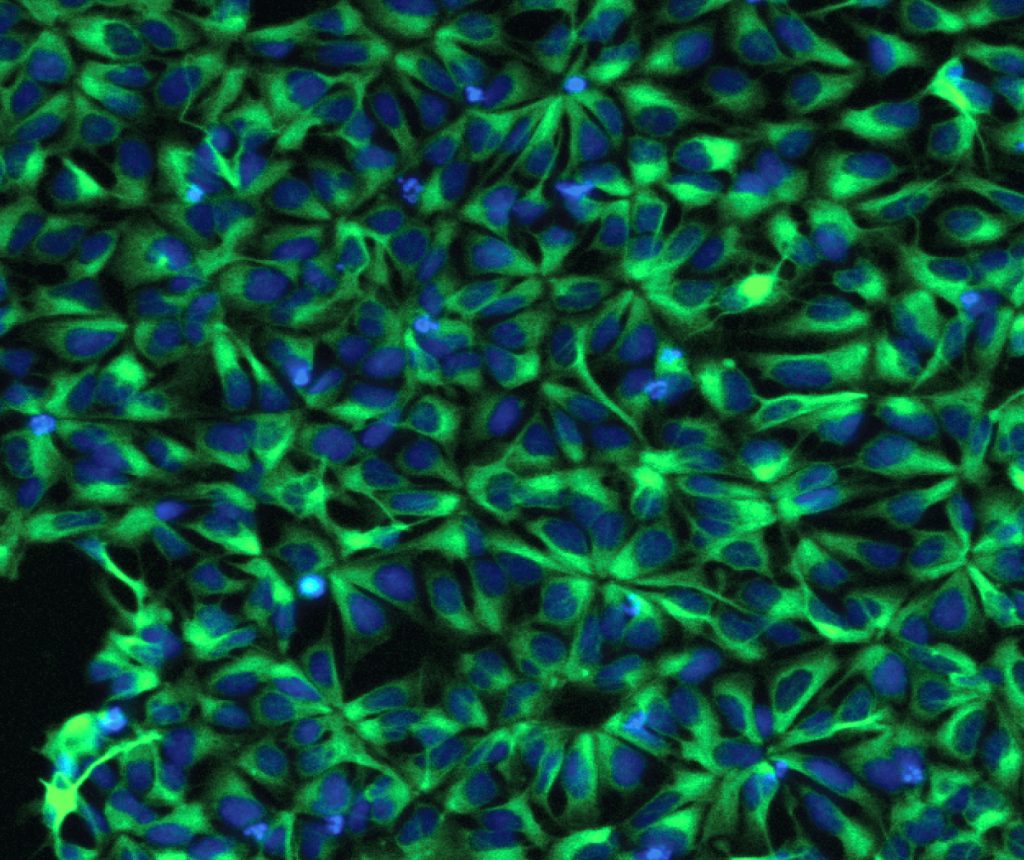
Human stem cell-derived neural progenitor cells. Credit Kristen Kroll.
Leaps and Bounds in the Study of Embryogenesis
In her early career, as a graduate student in the laboratory of Dr John Gerhart at the University of California in Berkeley, Dr Kroll developed a new method of transgenesis for the frog Xenopus laevis, a vertebrate animal model of embryonic development. Transgenesis involves introducing a gene into the genome of all cells of an organism.
The approach that Dr Kroll developed, in collaboration with Dr Enrique Amaya, has since been used by other researchers for many purposes, including analysing roles of particular genes in embryonic development and organ function. Transgenic lines of Xenopus generated by using this approach are in widespread use in the research community. Dr Kroll subsequently conducted postdoctoral work in the laboratory of Dr Marc Kirschner at Harvard Medical School, during which she identified a novel nuclear protein Geminin (Gmnn) and demonstrated roles for Gmnn in regulating Xenopus early embryonic development.
Following her graduate and postdoctoral training, Dr Kroll began a faculty position at Washington University School of Medicine. Work in her laboratory focused on studying how transcriptional and epigenetic regulation control embryonic development. Epigenetic regulation modulates gene expression without altering the underlying DNA sequence of the genome: DNA is compactly packed into a structure called chromatin, and modifying chromatin alters the accessibility or inaccessibility of parts of the genome, affecting the binding of transcription factors that regulate gene expression.
In early work in her laboratory, Dr Kroll and her colleagues demonstrated that Gmnn is a chromatin modifying protein essential for such epigenetic regulation to control multiple aspects of embryonic development. Her team demonstrated that embryonic cells require Gmnn to acquire a neural fate, as Gmnn promotes the expression of other genes that control aspects of later neural development, while Gmnn also antagonises acquisition of non-neural fates. This work defined several roles for Gmnn in normal embryonic development.
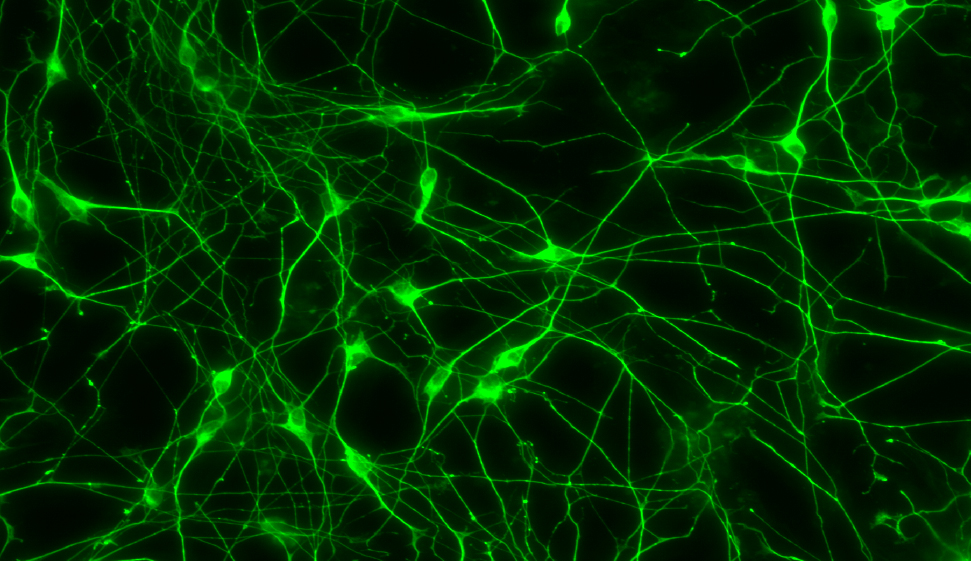
Human stem cell-derived interneurons. Credit Kristen Kroll.
Uncovering the Mechanisms of Neuron Development
As both the human genome sequence and many aspects of brain development are unique to humans, Dr Kroll and her team currently utilise stem cell-based approaches to model human neurodevelopment. They use directed differentiation of human pluripotent stem cell models (hPSCs) to identify regulatory networks that drive the development of particular neuronal cell types that are often disrupted to contribute to NDDs. Cortical interneurons (cINs) are one such cell type: these inhibitory neurons modulate the activity of excitatory neurons in the cerebral cortex. As excitatory and inhibitory activities in the cortex must remain balanced for normal cortical function, NDDs often stem from disruption of cIN development or function.
The majority of cINs are generated in a transient foetal brain structure located below the cortex called the medial ganglionic eminence (MGE). cINs generated in the MGE migrate into the cortex and establish connections with excitatory neurons to establish normal cortical neuron connectivity and function. To study cIN development, Dr Kroll’s team modified extant methods to enable the production of large numbers of MGE-like neural progenitors and MGE-derived cINs from hPSCs. This approach provides an experimentally accessible model for identifying mechanisms that control human cIN development. Subsequently, Dr Kroll’s team has focused on defining how gene expression is regulated by transcription factors and chromatin modifiers during this process.
One key regulator that they focused on was NKX2-1, a transcription factor required for cIN development. Their experiments revealed that one direct target of NKX2-1 is CHD2, a protein that remodels chromatin. Mutations in the CHD2 gene cause neurodevelopmental disorders including epilepsy, ASD, and intellectual disability. Research in the Kroll lab demonstrated that CHD2 deficiency disrupted cIN differentiation and altered cIN function. CHD2 and NKX2-1 co-bound genomic regulatory regions to co-regulate the expression of other genes required for cIN development. These studies helped identify important networks and key features of both normal and disrupted cIN development.
‘My lab focuses on defining regulatory networks controlling multiple aspects of neural development, identifying how epigenetic regulation modulates these networks, and determining how their dysregulation contributes to human neurodevelopmental disorders.’
Building New Models of Autism Spectrum Disorder
Dr Kroll’s team has also derived and characterised a number of new cellular models of ASD. The majority of ASD cases are polygenic (involving multiple genetic risk factors) and heritable, while most prior ASD cellular modelling efforts have focused instead on de novo (new rather than inherited) cases involving rare single-gene mutations. Therefore, Dr Kroll’s team attempted, for the first time, to model multiplex autism, the most common form of ASD involving inherited, polygenic risk, in patient-derived induced pluripotent stem cells (iPSCs). iPSC models were derived from three first-degree relatives with different degrees of inherited ASD (intermediate, severe, and clinically unaffected), and these were also compared with models from an unrelated, unaffected individual. iPSCs were differentiated into cINs and cortical excitatory neurons (cExNs), the major cell type that modulates cortical excitatory neuronal activity.
Cellular phenotyping showed that neural progenitor cells derived from ASD-affected family members exhibited increased cell death, by comparison with both models derived from unaffected relatives and an unrelated control. Both cExNs and cINs derived from individuals with ASD had distinct molecular signatures associated with the misregulated expression of genes associated with ASD, and those involved in neural development, function, behaviour, and cognition.
This novel study demonstrated that, even for ASD with complex aetiology, it is possible to detect affectation-linked cellular and molecular abnormalities, which can highlight pathways related to differing ASD affectation. Moreover, this work highlighted a need for the development of more cellular models that represent varied pathways to ASD, to define shared networks and pathways that are involved.
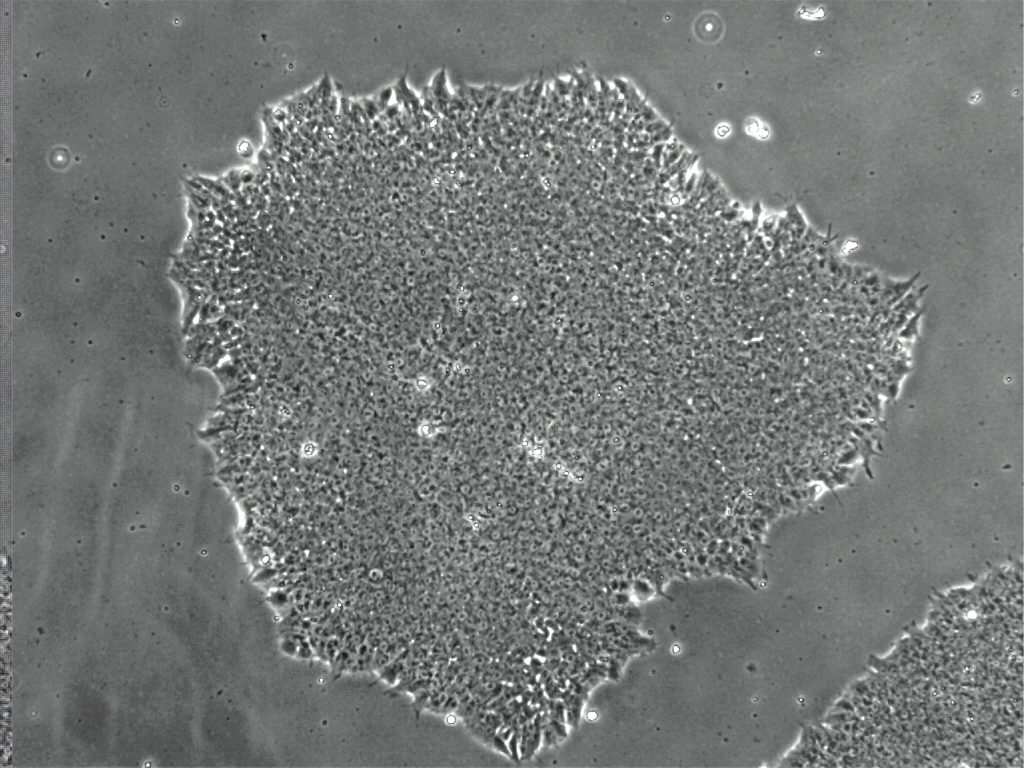
Human stem cell colony. Credit Kristen Kroll.
Determining Contributors to Clinical Phenotypes in Neuropsychiatric Disorders
Many neuropsychiatric disorders arise with variable penetrance (e.g., the extent to which carriers of a risk-related genetic variant express an associated phenotypic trait). One such example is copy number variants (CNVs) which micro-duplicate genomic intervals at chromosome 15q13.3, where the CHRNA7 channel gene is located. CNVs involving this gene cause severe clinical phenotypes in some individuals but not in others. This led Dr Kroll’s team to study iPSC models from a family carrying the same CNV, including duplication of only the CHRNA7 gene, but with different clinical phenotypes.
The team compared cExNs and cINs derived from two family members carrying the same CHRNA7 duplication, one of whom had an ASD diagnosis, while the other was clinically unaffected, as well as models from unrelated individuals without CHRNA7 duplication. Studying neurodevelopment and function in these models revealed that, while both affected and unaffected family members with CHRNA7 duplication had elevated CHRNA7 gene expression, models derived from the ASD affected individual had many neurodevelopmental phenotypes not observed in the other models. These included increased neuronal progenitor cell proliferation but defective neuronal differentiation, maturation, and migration, as well as increased endoplasmic reticulum stress involving failure to properly regulate cellular calcium trafficking.
Congruent with these findings, models from the ASD affected individual with CHRNA7 duplication exhibited reduced expression of genes involved in many neurodevelopmental processes. By contrast, the unaffected CHRNA7 duplication carrier instead exhibited upregulated expression of many genes in these same pathways, by comparison with controls lacking CHRNA7 duplication. These findings suggest that molecular compensation may have allowed normal neurodevelopment and an absence of clinical phenotypes in this unaffected individual, despite the presence of the same CNV at 15q13.3. Finally, as opposed to the neurodevelopmental phenotypes seen only in models from the affected individual, both models with CHRNA7 duplication exhibited altered neuronal function consistent with elevated channel activity.
This work indicates factors that may contribute to the typically variable phenotypic penetrance caused by CNVs involving genomic micro-duplication. Furthermore, the elevated endoplasmic reticulum stress and impaired neuronal migration seen in the model from the affected individual could be reversed by using pharmacological agents identified in this research. Therefore, these models may be useful for developing interventions to remedy diagnosis-associated anomalies, both for individuals with CHRNA7 duplication and for those with other copy number variants that cause similar phenotypes.
Current work in the Kroll laboratory focuses on using the types of modelling approaches described above to study a range of other neurodevelopmental disorders, in both subject-derived iPSC models and also in mouse models with the same mutation, particularly in cases involving the mutation of genes encoding transcription factors or chromatin modifiers.
Reference
https://doi.org/10.33548/SCIENTIA663
Meet the researcher

Dr Kristen L. Kroll
Washington University School of Medicine
Department of Developmental Biology
St. Louis, MO
USA
Dr Kristen Kroll is currently an Associate Professor of Developmental Biology at Washington University School of Medicine in St. Louis. Dr Kroll obtained her PhD from the University of California at Berkeley in 1994 and subsequently went on to complete a postdoctoral fellowship at Harvard Medical School in Boston. Dr Kroll’s research team focuses on the use of human pluripotent stem cell models to study gene regulatory networks that control neuronal development and to determine how disruption of these networks contributes to neurodevelopmental disorders. Dr Kroll has received multiple awards including the Damon Runyon-Walter Winchell Foundation Fellowship, March of Dimes Basil O’Connor Award, and American Cancer Society Hope Award, and has more than 50 publications in the field of developmental biology.
CONTACT
E: kkroll@wustl.edu
W: https://sites.wustl.edu/krolllab/
Twitter: https://twitter.com/krolllab
KEY COLLABORATORS
John Constantino, Washington University School of Medicine
James E. Huettner, Washington University School of Medicine
Bo Zhang, Washington University School of Medicine
Anita Bhattacharyya, Waisman Center, University of Wisconsin at Madison
Joseph Dougherty, Washington University School of Medicine
Lilianna Solnica-Krezel, Washington University School of Medicine
Thorold Theunissen, Washington University School of Medicine
CURRENT FUNDING
National Institute of Neurological Disorders and Stroke (National Institutes of Health)
National Institute of Mental Health (National Institutes of Health)
NIH Common Fund (National Institutes of Health)
Eunice Kennedy Shriver National Institute of Child Health and Human Development (National Institutes of Health)
Children’s Discovery Institute
M-CM Network
Jakob Gene Fund
FURTHER READING
EMA Lewis, K Meganathan, D Baldridge, P Gontarz, B Zhang, A Bonni, JN Constantino, KL Kroll, Cellular and molecular characterization of multiplex autism in human induced pluripotent stem cell-derived neurons, Molecular Autism, 2019, 10, 51, doi:10.1186/s13229-019-0306-0, PMCID: PMC6936127.
EMA Lewis, KL Kroll, Development and disease in a dish: the epigenetics of neurodevelopmental disorders, Epigenomics, 2018, 10, 219–231, doi:10.2217/epi-2017-0113, PMCID: PMC5810842.
K Meganathan, EMA Lewis, P Gontarz, S Liu, EG Stanley, AG Elefanty, JE Huettner, B Zhang, KL Kroll, Regulatory networks specifying cortical interneurons from human embryonic stem cells reveal roles for CHD2 in interneuron development, Proceedings of the National Academy of Sciences, 2017, 114, E11180–E11189, doi:10.1073/pnas.1712365115, PMCID: PMC5748186.
Want to republish our articles?
We encourage all formats of sharing and republishing of our articles. Whether you want to host on your website, publication or blog, we welcome this. Find out more
Creative Commons Licence
(CC BY 4.0)
This work is licensed under a Creative Commons Attribution 4.0 International License. 
What does this mean?
Share: You can copy and redistribute the material in any medium or format
Adapt: You can change, and build upon the material for any purpose, even commercially.
Credit: You must give appropriate credit, provide a link to the license, and indicate if changes were made.
More articles you may like
Laura Daniela Muñoz Duarte – Jo Philips | Rethinking Hydrogen Use in Acetogenic Bacteria: Kinetics, Thresholds, and Biotechnological Potential
Acetogenic bacteria can convert carbon dioxide and hydrogen into useful products, making them promising tools for sustainable biotechnology. Research led by Laura Muñoz and Jo Philips reveals that hydrogen consumption follows a simpler kinetic pattern than previously assumed, while also uncovering major differences between species in how efficiently they operate at low hydrogen levels. Together, these findings provide a clearer framework for selecting and optimising acetogens for industrial applications.
Dr Lynn George | A New Link Between Elongator Dysfunction and TDP-43 Loss in ALS
At Montana State University Billings, Dr Lynn George and her team are investigating the cellular mechanisms underlying amyotrophic lateral sclerosis (ALS). Using a new mouse model, their research reveals that disruption of the Elongator complex leads to nucleolar stress and the loss of the protein TDP-43 from motor neurons. These findings connect two major hallmarks of ALS and offer new insight into how the disease develops.
Dr David Hansen – Abhishek Juneja | Coming of Age: Active Balancing in Adolescence
Adolescence, the transitional stage between childhood and adulthood, comes with challenges as well opportunities for growth and learning.
Jella van de Laak | Could Tumour- Targeting Bacteria Enhance Cancer Immunotherapy?
Cancer immunotherapy has transformed treatment outcomes, yet many patients still fail to respond. At Maastricht University, Jella van de Laak, working with Professor Ludwig Dubois and Professor Jan Theys, is exploring how tumour-targeting bacteria could improve immune responses and expand treatment success.




