Professor Maria Ngu-Schwemlein – Developing Mercury Immobilisers and Antibiotic Enhancers
The rise of antibiotic-resistant strains of bacteria and the ubiquitous nature of mercury in the environment are two major health hazards and concerns faced today. Professor Maria Ngu-Schwemlein and her team from the Winston-Salem State University, USA, are addressing these issues by developing biomolecules that can bind and immobilise toxic mercury ions more effectively. Together with Professor Ngu-Schwemlein’s collaborators, the team is also investigating carbon nanodots as molecular scaffolds to enhance antibiotics in combating antimicrobial resistance.
Finding and Enveloping the Target
Mercury and many other heavy metals can be found in water as positively charged ions. Typical concentrations of metal ions in untreated water supplies are usually quite low and not all metals are harmful – indeed, some are vital. Some metal ions are required for a multitude of biochemical reactions taking place within living organisms while others, such as mercury, are toxic. Therefore, developing methods that can reliably target and remove mercury ions, over other metals, is a major undertaking.
Because of the health hazards posed by mercury, highlighted by the public health disasters in Iraq and Minamata Bay in Japan, many international efforts have been initiated to remediate contaminated sites, protect human health, and reduce mercury pollution. Although various technologies have been developed, there is still a need for compounds that can effectively bind to mercury – a process called ‘immobilisation’ – so that it can be more easily removed. Medical researchers are also constantly trying to develop new therapies that can target and effectively immobilise mercury ions so that they can be removed from patients.
In addition to the health hazards posed by mercury, the rise of antibiotic-resistant bacteria is another global challenge threatening human health. As a result, the World Health Organization (WHO) has endorsed a global action plan, which aims to not only prevent but treat infectious diseases with effective medicines that can target and kill harmful, antibiotic-resistant bacteria.
Professor Maria Ngu-Schwemlein and her team from the Winston-Salem State University are designing new compounds that tackle both of these challenges. In one of their research projects, they have modified and improved existing molecules that can target mercury ions for safe remediation. In their second research direction, the team has also devoted time to find compounds that can enhance the effect of antibiotics on resistant strains of bacteria.
The mechanism by which each type of compound binds to a metal ion or a bacterium is similar. Take, for example, an insect-eating sundew plant; this species is an effective carnivore partly because its tentacles can envelope their prey. Each point of contact is comparatively weak, but in combination, they provide a very firm grip on their target. This behaviour is somewhat analogous to the binding action of the molecules designed by Professor Ngu-Schwemlein and her colleagues. In general, the team first identifies which atom or group of atoms bind to their target. They then start modifying the surrounding structure in order to maximise the number of points of contact or bonds formed.

Caged Mercury Ions
Sulphur atoms have a particularly high affinity for mercury ions. Some sulphur-containing compounds, including those found naturally in proteins, comprise not one, but several sulphur atoms. This means that each compound can form multiple contact points or bonds with a single mercury ion. There are other atoms, such as oxygen and nitrogen, that can also bind to mercury, though this tends to result in a relatively weak interaction.
The sulphur-containing compound that binds to a mercury ion with two or more contact points is an example of a ‘chelator’. One of the goals of Professor Ngu-Schwemlein’s research is to develop a chelator that can wrap around a mercury ion. The wrapping of mercury immobilises it and prevents it from interacting with other biological molecules. In principle, this approach should protect the body from the toxic effects of mercury. While there is an understanding about how chelators behave, the team believed that more could be done to build more effective mercury deactivators.
As part of a preliminary investigation, Professor Ngu-Schwemlein’s group determined how sulphur-containing units in a compound could bind mercury ions. By using a technique called ‘electrospray ionisation mass spectrometry’ to analyse these interactions, they found that tethering multiple sulphur-containing units to a chelator compound can enhance the binding efficiency through the compound’s ability to bind more than one mercury ion. The compound that forms when a chelator is bound to a metal ion is known as an ‘adduct’.
For the next phase of the study, the team was interested in finding out how the ratios of the chelated mercury ions would change with time. In other words, do they convert from one type to another, and if so, how does this affect the strength of the interaction? In addition, the team wanted to explore the effect of incorporating other atoms that could bind to mercury.
The researchers narrowed their study to a class of naturally occurring peptide chelators that contain a sulphur group at each end. The chelator is symbolised here as SXS, where X is a group of atoms that connect the two sulphur groups (S), which can bind to mercury. The team prepared two types of SXS compounds, one in which X contains oxygen atoms that could also bind to mercury, and another variant where X is not expected to bind to mercury.
After multiple experiments, the team found that both variants bound to mercury in a ratio of 1:1. The team also found that the presence of the oxygen atoms at X appeared to speed up the formation of the adduct. This shows that the oxygen atoms enhanced the interaction between the chelator SXS and the mercury ion. More specifically, the oxygen atoms formed a type of bond with mercury, which helped to bring the chelator closer and more tightly to mercury.
In collaboration with Dr John Merle, the group also performed computational studies. Their calculations and simulations also confirmed that the extra oxygen atoms would result in a stronger, more tightly formed adduct.
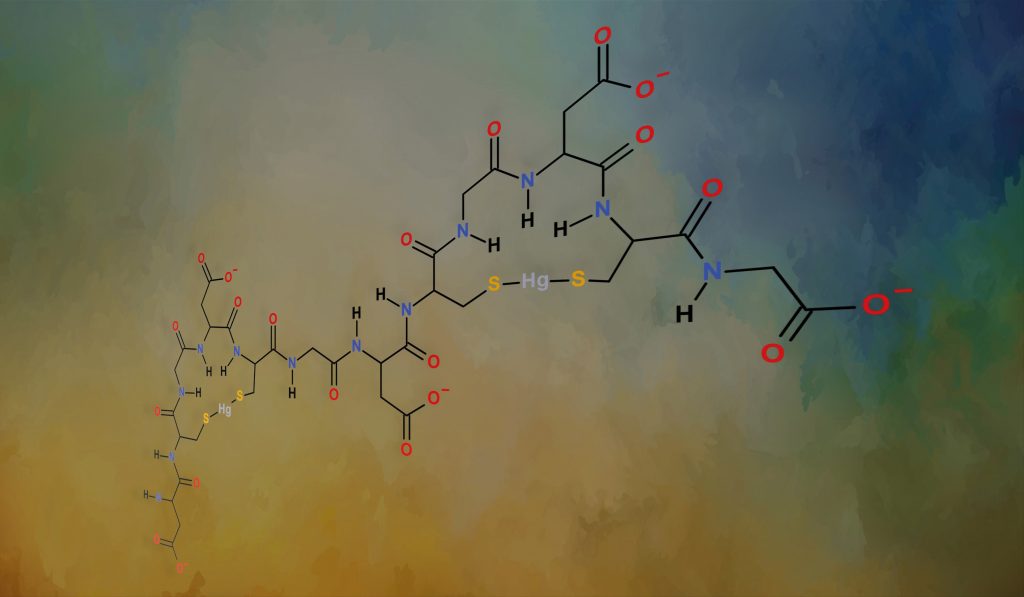
Indole Rings and Spatial Orientation
Following the success of the above investigation, Professor Ngu-Schwemlein and her team turned their attention to studying more structurally diverse groups of atoms at X and how these groups influence the chelation of mercury. The team wanted to learn more about how a fused network of atoms, known as an ‘indole ring’, would shield the chelated mercury ion from reacting with other molecules. One can visualise this interaction by holding a tennis ball above the face of a tennis racket. The racket frame represents the indole ring, and the tennis ball represents the mercury ion. Indoles have an appreciable amount of negative electron density and are attracted to positively charged mercury ions. When an indole ring is bound to a mercury ion, it forms a barrier on one side of the mercury ion.
The researchers compared two variants of SXS, both of which contained an indole ring. The only difference between the variants was the direction in which the rings were pointing. Interestingly, the team confirmed that only one of the indole-containing variants, with the ring pointing towards the mercury ion, interacted strongly. The specific orientation of the indole rings was also supported by computational studies. The researchers are currently working on incorporating two indole rings into the variants to ‘sandwich’ and shield the chelated mercury ion, thereby creating a barrier on both sides of the mercury ion. Such chelators may provide a valuable tool for immobilising toxic mercury ions more effectively. They hope that these findings will be useful for applications including environmental remediation, and human health protection.

Combatting Antibiotic-Resistant Bacteria
Another major part of Professor Ngu-Schwemlein’s research is the development of compounds that combat antibiotic-resistant bacteria, with the assistance of compounds known as ‘dendrimers’. Dendrimers can be viewed as compounds where identical branches of atoms extend from a central point, like the tentacles of an octopus. What makes their selected dendrimers behave similarly to chelators, is that along each branch of the dendrimer are positively charged nitrogen that can, under appropriate conditions, bind to the cell walls of bacteria. When this happens, the cell walls are weakened and, in a few cases, broken down, killing the bacteria.
The effectiveness of dendrimers as antibacterial compounds is limited, and this is thought to be due to the low concentration of nitrogen groups. Consequently, the team wanted to find a way of packing more nitrogen groups into a small space, thereby increasing the chances of disrupting the cell walls on contact.
In order to pack more of these groups into a smaller space, Professor Ngu-Schwemlein and her collaborator Dr Suk-Fun Chin utilised a class of materials known as ‘carbon nanodots’. As the name implies, carbon nanodots are small particles of carbon. The team chose carbon nanodots derived from starch as a central framework, because they have the surface properties needed to attach multiple dendrimers.
Once the nanodots were grafted with dendrimers, the team targeted two common types of bacteria, E. Coli and S. aureus, as well as three other bacteria that have evolved resistance to antibiotics. The team was pleased to discover that these dendrimer-bound carbon nanodots were effective at killing all types of bacteria.
Given that the team’s dendrimer-bound nanodots were able to disrupt the cell walls of bacteria, the researchers then decided to combine them with some existing common antibiotics. In many antibiotic-resistant bacteria, the cell walls form a barrier to these common antibiotics. Much to their delight, the researchers found that the antibiotics were indeed made more effective when administered with their nanodots.
These results are highly significant. It shows that there are other treatments, well within reach, that can be utilised to combat antibiotic-resistant bacteria. Professor Ngu-Schwemlein and her team hope that their findings will encourage others to build on their success.
Reference
https://doi.org/10.33548/SCIENTIA653
Meet the researcher

Professor Maria Ngu-Schwemlein
Department of Chemistry
Winston-Salem State University
Winston-Salem, NC
USA
Professor Maria Ngu-Schwemlein was awarded her PhD from the Australian National University in 1989. After working at many notable institutes, including ETH Zürich, Louisiana State University and Lawrence Livermore National Laboratory, she joined Winston-Salem State University in 2006 as an Associate Professor, before becoming a full Professor in 2010. Her recent research interests focus on designing second-generation chelators for toxic metal ions and developing antimicrobials to combat antibiotic-resistant bacteria. Professor Ngu-Schwemlein is passionate about training undergraduates in research skills, and she involves them in her projects, providing them with invaluable research experience. In recognition of her contributions at Winston-Salem State University, Prof Ngu-Schwemlein was awarded the Wilveria Atkinson Distinguished Research Award and the John Fountain Master Teaching Award.
CONTACT
E: schwemleinmn@wssu.edu
W: https://www.wssu.edu/profiles/schwemleinmn/index.html
KEY COLLABORATORS
Dr Suk Fun Chin, Universiti Malaysia Sarawak
Dr XiuLi Lin, Wake Forest University
Dr John Merle, Winston-Salem State University
FUNDING
US National Science Foundation
The National Institutes of Health
FURTHER READING
M Ngu-Schwemlein, J Merle, W Meeker, K Risdon-Langdon, T Nixon, Evaluating the involvement of tryptophan on thiolated peptide-mercury(II) complexes: Cation-pi interactions, Inorganica Chimica Acta, 2020, 506, 119552.
J Merle, J Mazlo, J Watts, R Moreno, M Ngu-Schwemlein, Reaction mixture analysis by ESI-MS: Mercury(II) and dicysteinyl tripeptide complex formation, International Journal of Mass Spectrometry, 2018, 426, 38.
M Ngu-Schwemlein, X Lin, B Rudd, M Bronson, Synthesis and ESI mass spectrometric analysis of the association of mercury(II) with multi-cysteinyl peptides, Journal of Inorganic Biochemistry, 2014, 133, 8.
M Ngu-Schwemlein, SF Chin, R Hileman, C Drozdowski, C Upchurch, A Hargrove, Carbon nanodots as molecular scaffolds for development of antimicrobial agents, Bioorganic & Medicinal Chemistry Letters, 2016, 26, 1745.

Want to republish our articles?
We encourage all formats of sharing and republishing of our articles. Whether you want to host on your website, publication or blog, we welcome this. Find out more
Creative Commons Licence
(CC BY 4.0)
This work is licensed under a Creative Commons Attribution 4.0 International License. 
What does this mean?
Share: You can copy and redistribute the material in any medium or format
Adapt: You can change, and build upon the material for any purpose, even commercially.
Credit: You must give appropriate credit, provide a link to the license, and indicate if changes were made.
More articles you may like
Grandmothers: Innovation Through Tradition
Grandmother Project – Change through Culture (GMP) is an organisation dedicated to documenting the role of grandmothers and demonstrating the effectiveness of grandmother-inclusive strategies in improving the health and well-being of women, children, and adolescents. GMP’s groundbreaking work challenges conventional wisdom to transform community-based interventions in Africa and beyond, harnessing a powerful but often overlooked resource: the wisdom and influence of grandmothers.
Dr Robert Larkin | Cultivating Change to Improve Soil Health and Increase Potato Yield
Environmental quality and food production are facing the pressing challenges of climate change and global population growth. Dr Robert Larkin from the United States Department of Agriculture-Agricultural Research Service (USDA-ARS) and a team of plant scientists developed and tested a range of crop management systems to help overcome these compounding challenges. Their work is improving soil health and increasing the yield of potato crops, contributing to the future food security of nations.
Professor Giorgio Buttazzo | Artificial Intelligence and a Crossroads for Humanity
Where do we stand with artificial intelligence? Might machines take over our jobs? Can machines become conscious? Might we be harmed by robots? What is the future of humanity? Professor Giorgio Buttazzo of Scuola Superiore Sant’Anna is an expert in artificial intelligence and neural networks. In a recent publication, he provides considered insights into some of the most pressing questions surrounding artificial intelligence and humanity.
Dr Ralf Adam | New Technologies Shaping the Future of Oral Hygiene
Understanding the efficiency of various toothbrush technologies is essential for achieving optimal oral health. Dr Ralf Adam, who leads a dedicated team at Procter & Gamble in Germany, is keen to investigate the complexities of these technologies. His team have provided new insights into the best toothbrush types for plaque removal and the maintenance of gum health. By highlighting the importance of informed oral care decisions and ongoing investigations, this vital research works towards ensuring everyone can achieve a brighter, healthier smile.




