Professor Michael O’Donnell – A Twin DNA Replication Factory
For life on Earth to grow, its genetic material must be copied and reproduced in a process known as DNA replication. Professor Michael O’Donnell, head of the Rockefeller University’s DNA replication laboratory, has devoted his over 30-year career to the study of the protein complex that is responsible for just that – the replisome. Recently, Professor O’Donnell and his team uncovered exciting insights into the function of this remarkable piece of molecular machinery.
Behind the Scenes of DNA Replication
From conception, a single fertilised egg cell faces exponential growth in order to become a two trillion cell new-born. During each round of cell division, DNA – copied at a rate of 25 nucleotide units per second – requires extreme accuracy during replication, since any mistake has the potential to be fatal. DNA stores the genetic blueprint of every organism in a series of nucleotide chemical bases known as adenine (A), thymine (T), guanine (G), and cytosine (C). These four letters form a unique genetic code underpinning the characteristics of nearly every life form on Earth.
DNA consists of two complementary, anti-parallel strands that coil around a common axis in the shape of a double helix. A base from one strand pairs with a base from the anti-parallel strand in the following manner: A pairs with T, and G pairs with C. While the order of bases is ever-changing, the manner in which they pair is not. It may seem paradoxical, therefore, that DNA replication occurs so smoothly. Enter the replisome.
‘Every time that a cell divides to form two new cells, the DNA instructions for life must be duplicated in a timely and accurate fashion’, emphasises Professor Michael O’Donnell from the Rockefeller University, describing what has been the focus of his work for over thirty years. ‘This remarkable feat is accomplished by a machinery somewhat like a sewing machine composed of many protein “gears” that function together, referred to as a replisome.’
Professor O’Donnell heads the Rockefeller University’s DNA replication laboratory and has devoted his career to understanding the unique architecture of proteins contained within the replisome, by studying both their physical structures and biochemical activities. His work aims to provide new insights into cellular replication, repair, and genetic inheritance.
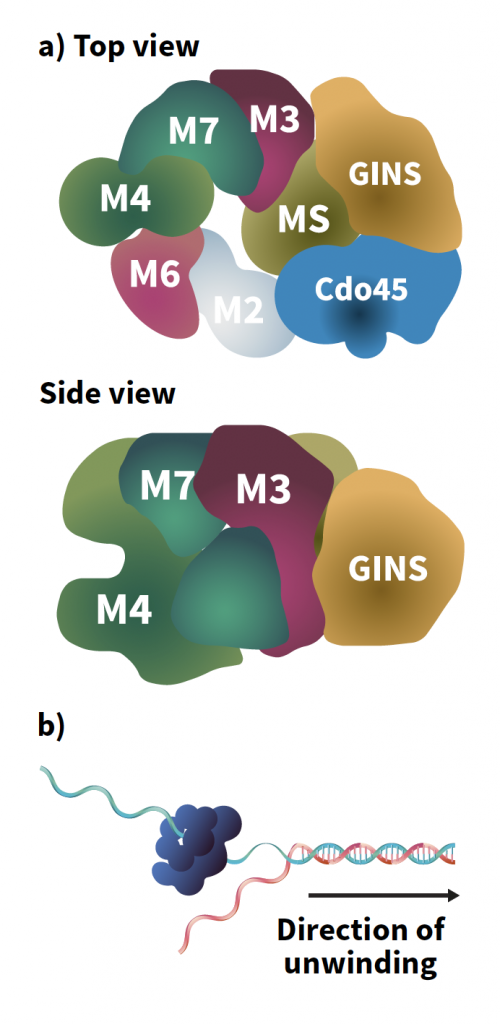
Figure 1: CMG helicase. A) Top and side views of CMG, top, side. B) DNA unwinding is achieved by helicase encircling one strand and excluding the other strand. The helicase motors along the strand it encircles and acts like a wedge to split the DNA duplex into two single-strands.
From Baker’s Yeast to Humans
Over a billion years of evolution, the components of the replisome have been largely conserved across the animal, plant and fungi kingdoms. One of the leading protein ‘gears’ found within is helicase, responsible for separating the two DNA strands. DNA must be unwound for replication to occur, going from a tightly coiled double helix to two straight lines, much like the unzipping of a zip. From baker’s yeast to humans, the helicase found within the replisome is an 11-protein assembly called CMG, named for its three components: Cdc45 protein, the MCM2-7 motor ring, and GINS complex.
This unwinding is achieved by CMG encircling one strand of a double-stranded DNA molecule via its MCM ring and excluding the other strand to the outside of the ring. Then, CMG tracks along the encircled strand, acting as a moving wedge to split the DNA duplex apart.
CMG’s other two elements – the Cdc45 protein and GINS complex – are used to attract and capture other protein components of the replisome machine. These include DNA polymerases, used to synthesise DNA by assembling individual nucleotides (building blocks) of DNA; a ring-shaped sliding clamp called PCNA, used to tether DNA polymerases to parent strands of DNA; a clamp loader protein that allows PCNA to clamp around DNA; and a primase, called DNA polymerase α-primase (DNA pol α-primase), which synthesises primers required to initiate DNA replication.
‘Every time that a cell divides to form two new cells, the DNA instructions for life must be duplicated in a timely and accurate fashion…This remarkable feat is accomplished by a machinery somewhat like a sewing machine composed of many protein “gears” that function together, referred to as a “replisome”’.
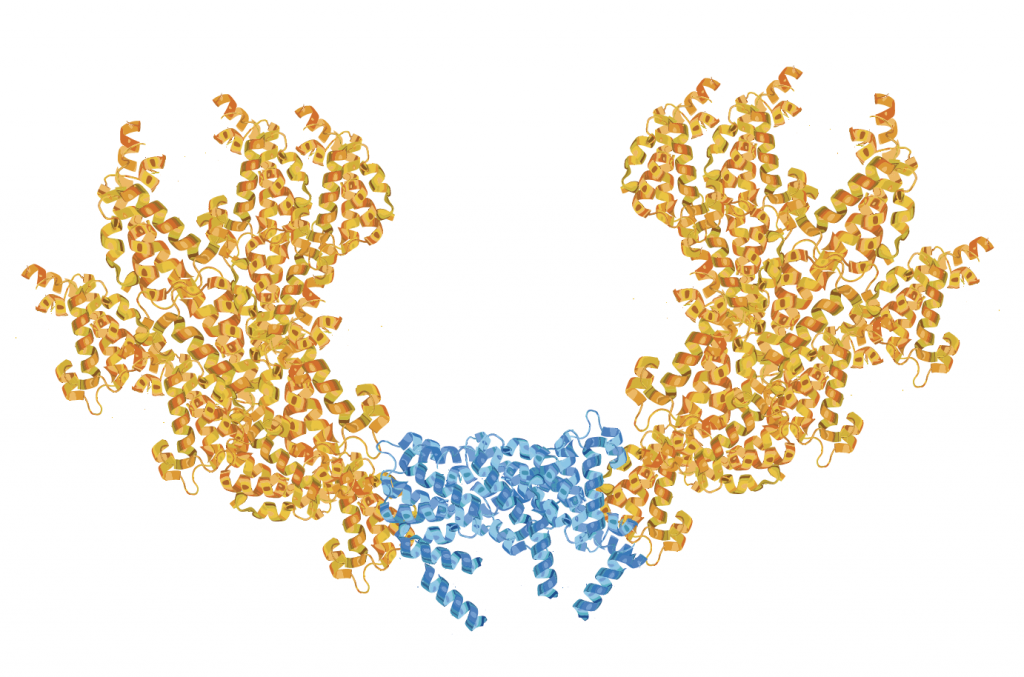
Figure 2. Twin CMGs. The Ctf4 trimer (blue) holds two CMG helicases (orange) in a nearly vertical position relative to the Ctf4 disk. This structure is based on Z Yuan et al, 2019. The Ctf4 trimer organises two sister replisomes and one primase-polymerase alpha into a replication factory core.
A Factory of Twin Replisomes
DNA replication occurs in the nucleus, or central organelle, of each cell. ‘It has long been known that replication proteins localise to various “spots” in the nucleus that can be observed in the microscope and are referred to as replication foci’, explains Professor O’Donnell. ‘These are the sites of DNA replication. Studies in yeast showed that these spots mainly consist of only two replisomes. Therefore, it would appear that replication occurs in factories of twin replisomes.’
Indeed, Professor O’Donnell and colleagues discovered that a novel player is crucial to the formation of the twin replisome factory: a protein called Ctf4 (Chromosome Transmission Fidelity 4). Ctf4 is a homotrimer, meaning it has three identical subunits, and two of these are used to tightly bind CMG molecules. Nevertheless, the exact structure of this twin CMG-Ctf4 unit remained unknown. This is where Professor O’Donnell teamed up with Dr Huilin Li of the Van Andel Institute in Michigan, an expert in a technique known as cryogenic electron microscopy (cryo-EM) used to examine the high-resolution structures of biomolecules.
With the help of Dr Li, Professor O’Donnell discovered that the mystery CMG-Ctf4 structure was that of two CMG helicases oriented in a head-to-head fashion around one Ctf4 protein. This unique layout explains the observation that DNA foci in yeast contain two replisomes. In humans, however, nuclear foci are bigger but super high-resolution imagery has revealed that each large focus is simply composed of many sub-foci, and each of these sub-foci is a twin replisome. Professor O’Donnell and his team thus uncovered a key finding underpinning the DNA replication factory: twin replisomes are held by a Ctf4 scaffold in a head-to-head fashion.
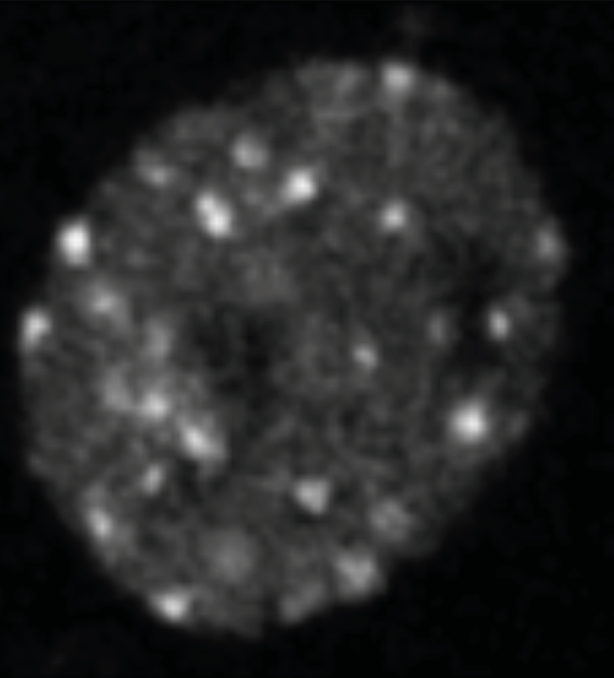
Figure 3: Nuclear foci. PCNA was fused to GFP and imaged in yeast nuclei. Citation: E Kitamura, et al, 2006. Live-cell imaging reveals replication of individual replicons in eukaryotic replication factories, Cell, 125, 1297–1308.
CMG Unwinds DNA in the N-first Direction
DNA replication is an inherently dynamic process: a helicase must traverse an entire DNA molecule in order to split it into two strands and enable duplication. CMG, like other proteins, has an asymmetric structure, defined by the N- and C-ends of the proteins. The orientation of CMG as it travels on DNA was another mystery that the O’Donnell laboratory was keen to solve. Does it travel N-first or C-first?
The replication factory mechanism occurs as follows: a parental, double-stranded DNA molecule feeds into the system and is separated into two single strands. Of these, one strand threads through CMG, while the other remains outside the CMG ring at the centre of the factory. Professor O’Donnell, with the help of Dr Li, used cryo-EM to enable direct visualisation of this process to investigate CMG’s direction of travel. ‘We determined the orientation of CMG helicase while it travels on DNA, which was opposite from the orientation that had been assumed by the field for over a decade. This “challenge” has been confirmed by other labs now’, explains Professor O’Donnell.
This visualisation showed the fates of each DNA strand. After being threaded through CMG, one DNA strand was duplicated by a polymerase ε protein (DNA pol ε). DNA pol ε, which directly bound CMG, was held in place by the aforementioned PCNA clamp which encircles double-stranded DNA. The duplication of the strand not threaded through CMG, termed the lagging strand, was attended by DNA polymerase δ (DNA pol δ). DNA pol δ replicated the lagging strand thanks to the primers synthesised by DNA pol α-primase while also being held in place by the PCNA sliding clamp.
This cryo-EM visualisation led to another key discovery in solving the replisome puzzle: only one DNA pol α-primase could bind the Ctf4 trimer. Consisting of three subunits, the Ctf4 trimer, therefore, bound two of these to CMG molecules, and the third to DNA pol α-primase, implying that DNA pol α-primase must split its primer forming activity between the two lagging strands. Slowly but surely, Professor O’Donnell was shining a light on the previously hidden intricacies of the replisome.
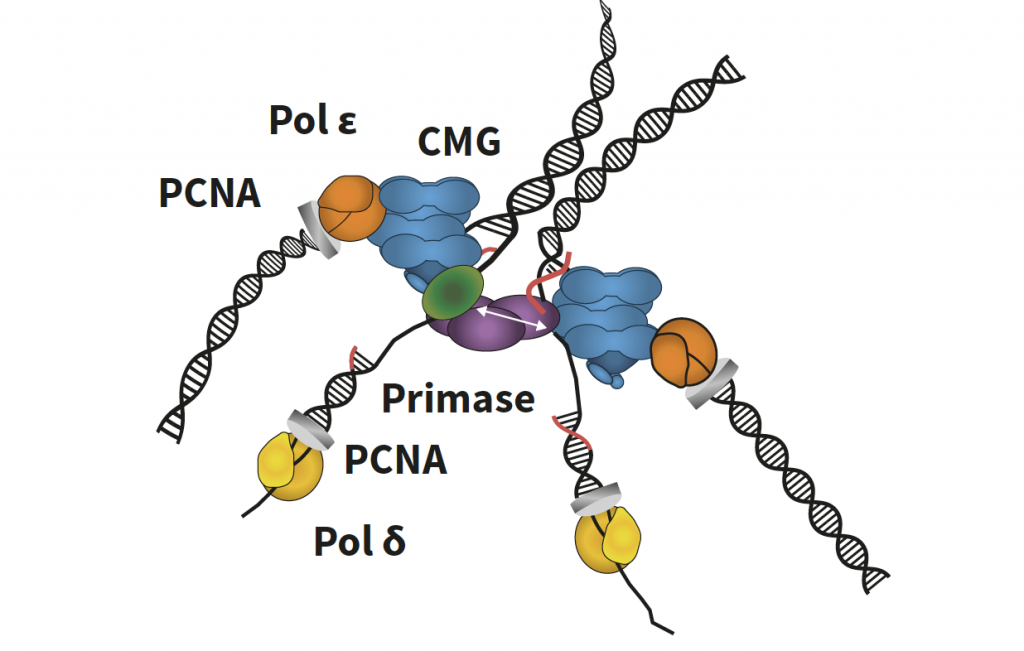
Figure 4. Twin replisome factory. Twin replisomes held by a Ctf4 scaffold. Two CMGs (blue) are held by one Ctf4 trimer (purple). The original parental DNA duplex feeds into each CMG from the centre of the factory and is split at the surface, where one strand goes through CMG, where it is duplicated by Polymerase epsilon (orange)-PCNA clamp (grey) upon its exit from CMG. One flexible primase (green) binds the third Ctf4 subunit and alternates activity between the two strands to form primers (red) that are extended by Polymerase delta (yellow) held to DNA by the PCNA clamp (grey).
An Ensured Inheritance
Perhaps surprisingly, every bodily cell contains the entire selection of DNA or genome. However, only certain genes end up being ‘switched on’. Whether or not a gene is expressed depends on epigenetic markers, which, in a nutshell, are chemical tags on DNA. DNA is packaged inside a cell into functional units called nucleosomes which include these markers, ready to determine the function of each cell. Thus, epigenetic inheritance is an essential component of cell division which must be preserved by the replisome. In the model described by Professor O’Donnell, two CMGs bind to one nucleosome, suggesting that the twin replication factory itself facilitates the transfer of nucleosomes to new DNA, ensuring developmental epigenetic inheritance is achieved.
Professor O’Donnell and his team also proposed that the replication factory helps to organise the newly synthesised DNA genome and that the twin model is able to communicate with itself. Professor O’Donnell summarised that ‘it is possible the two replisomes communicate their status to one another, such that if one replisome stops due to DNA damage, the other replisome may stop as well’. This communication could be integral for the production of healthy daughter cells, void of rogue DNA damage that could lead to cancer.
Looking to the Future
Professor O’Donnell’s laboratory is looking to validate the conclusions drawn here, both in live mammalian cell studies and in bacteria. Presumably, the twin replication factory is applicable to all life forms and provides key information for furthering the study of DNA replication. The enthusiasm that Professor O’Donnell shares for unlocking new secrets of this field is contagious, as he declares ‘the current study is only the beginning of a comprehensive understanding of how replication is organised in the cell. We expect that additional proteins, further layers of organisation, and yet to be determined dynamic actions of these proteins, exist in nuclear replication factories.’
Reference
https://doi.org/10.33548/SCIENTIA546
Meet the researcher

Professor Michael O’Donnell
Department of DNA Replication
The Rockefeller University
New York, NY
USA
Professor Michael O’Donnell received his PhD in biochemistry from the University of Michigan in 1982 and went on to complete a postdoctoral position at Stanford University in 1986. At the close of his postdoctoral studies in 1986 he founded his research group at Cornell Medical College in New York City, which he moved to the Rockefeller University in 1996 where he now heads the Laboratory of DNA Replication. Over his 30-year career, he has achieved an impressive accolade of honours and awards including becoming an investigator with the Howard Hughes Medical Institute in 1990 to the present time, and induction into the United States National Academy of Sciences in 2006. Alongside this, he is an editor and reviewer for several research journals and fundraising bodies. His research focuses on mechanistically understanding how the collection of proteins involved in DNA replication ensures genomic integrity. During the COVID-19 outbreak, Professor O’Donnell and his team are investigating potential vulnerabilities in the reproduction of coronavirus.
CONTACT
E: odonnel@mail.rockefeller.edu
W: https://www.rockefeller.edu/our-scientists/heads-of-laboratories/877-michael-odonnell/
https://www.hhmi.org/scientists/michael-e-odonnell
http://odonnell.rockefeller.edu
KEY COLLABORATORS
Dr Huilin Li, Van Andel Institute, Grand Rapids, Michigan
Dr John Kuriyan, University of California at Berkeley, Berkeley, California
FUNDING
National Institute of Health General Medical Institute
Howard Hughes Medical Institute
FURTHER READING
Z Yuan, R Georgescu, R Santos, D Zhang, L Bai, G Zhao, M O’Donnell, H Li, Ctf4 organizes sister replisomes and Pol α into a replication factory, eLife, 2019, 8, e47405.
R Georgescu, Z Yuan, L Bai, R Santos, J Sun, D Zhang, O Yurieva, H Li, M O’Donnell, Structure of eukaryotic CMG helicase at a replication fork and implications to replisome architecture and origin initiation, Proceedings of the National Academy of Sciences, 2017, 114, 5, E697–E706.
Z Yuan, L Bai, J Sun, R Georgescu, J Liu, M O’Donnell, H Li, Structure of the eukaryotic replicative CMG helicase suggests a pumpjack motion for translocation, Nature Structural & Molecular Biology, 2016, 23, 3, 217–224.

Want to republish our articles?
We encourage all formats of sharing and republishing of our articles. Whether you want to host on your website, publication or blog, we welcome this. Find out more
Creative Commons Licence
(CC BY 4.0)
This work is licensed under a Creative Commons Attribution 4.0 International License. 
What does this mean?
Share: You can copy and redistribute the material in any medium or format
Adapt: You can change, and build upon the material for any purpose, even commercially.
Credit: You must give appropriate credit, provide a link to the license, and indicate if changes were made.
More articles you may like
Grandmothers: Innovation Through Tradition
Grandmother Project – Change through Culture (GMP) is an organisation dedicated to documenting the role of grandmothers and demonstrating the effectiveness of grandmother-inclusive strategies in improving the health and well-being of women, children, and adolescents. GMP’s groundbreaking work challenges conventional wisdom to transform community-based interventions in Africa and beyond, harnessing a powerful but often overlooked resource: the wisdom and influence of grandmothers.
Dr Robert Larkin | Cultivating Change to Improve Soil Health and Increase Potato Yield
Environmental quality and food production are facing the pressing challenges of climate change and global population growth. Dr Robert Larkin from the United States Department of Agriculture-Agricultural Research Service (USDA-ARS) and a team of plant scientists developed and tested a range of crop management systems to help overcome these compounding challenges. Their work is improving soil health and increasing the yield of potato crops, contributing to the future food security of nations.
Professor Giorgio Buttazzo | Artificial Intelligence and a Crossroads for Humanity
Where do we stand with artificial intelligence? Might machines take over our jobs? Can machines become conscious? Might we be harmed by robots? What is the future of humanity? Professor Giorgio Buttazzo of Scuola Superiore Sant’Anna is an expert in artificial intelligence and neural networks. In a recent publication, he provides considered insights into some of the most pressing questions surrounding artificial intelligence and humanity.
Dr Ralf Adam | New Technologies Shaping the Future of Oral Hygiene
Understanding the efficiency of various toothbrush technologies is essential for achieving optimal oral health. Dr Ralf Adam, who leads a dedicated team at Procter & Gamble in Germany, is keen to investigate the complexities of these technologies. His team have provided new insights into the best toothbrush types for plaque removal and the maintenance of gum health. By highlighting the importance of informed oral care decisions and ongoing investigations, this vital research works towards ensuring everyone can achieve a brighter, healthier smile.




