Dr Jin Yu – Remdesivir: Halting the Viral Replication of SARS-CoV-2
The design of effective antivirals is a key priority in the global effort to curb the pandemic caused by the novel coronavirus SARS-CoV-2. The FDA-approved drug Remdesivir (RDV) acts by interfering with the SARS-CoV-2 viral replication mechanism. Dr Jin Yu and her team from the University of California, Irvine, conducted a computational study to elucidate how the RNA-dependent RNA polymerase (RdRp), responsible for SARS-CoV-2 genomic replication, is inhibited by RDV. Excitingly, they showed that RDV binds tightly to RdRp, stabilising a closed conformation of the active site for successful incorporation to effectively halt viral replication.
COVID-19: A Rapidly Evolving Disease
Almost two years on from the start of the COVID-19 pandemic, there have been more than 240 million confirmed cases of the disease and almost 5 million deaths linked to COVID-19 worldwide. According to the World Health Organization, COVID-19 remains a rapidly evolving disease that continues to push healthcare systems beyond their current capacities in most countries across the globe.
As the SARS-CoV-2 virus continues to circulate, more variants will continue to emerge. To date, four variants have been designated specific variants of concern, all of which are even more transmissible than the ancestral strain of SARS-CoV-2. Currently, a further five variants of interest are being closely monitored and evaluated. Alongside the implementation of a global vaccination effort, the development of effective antiviral treatments targeting SARS-CoV-2 is a primary focus in the global effort to curb the COVID-19 pandemic.
Using a Nucleotide Analogue to Jam the RNA Synthesis Mechanism
SARS-CoV-2 is a newly emerged member of the family of coronaviruses. Coronaviruses are enveloped, which means they contain an outermost layer that protects their genetic material when travelling between host cells. SARS-CoV-2 is also a single-stranded ribonucleic acid (RNA) virus. RNA is a multifunctional molecule that, among other capabilities, works to convert genetic information into proteins.
The key RNA synthesising engine of the replication-transcription machinery encoded in the genomes of all RNA viruses, including SARS-CoV-2, is the RNA-dependent RNA polymerase (RdRp) enzyme.
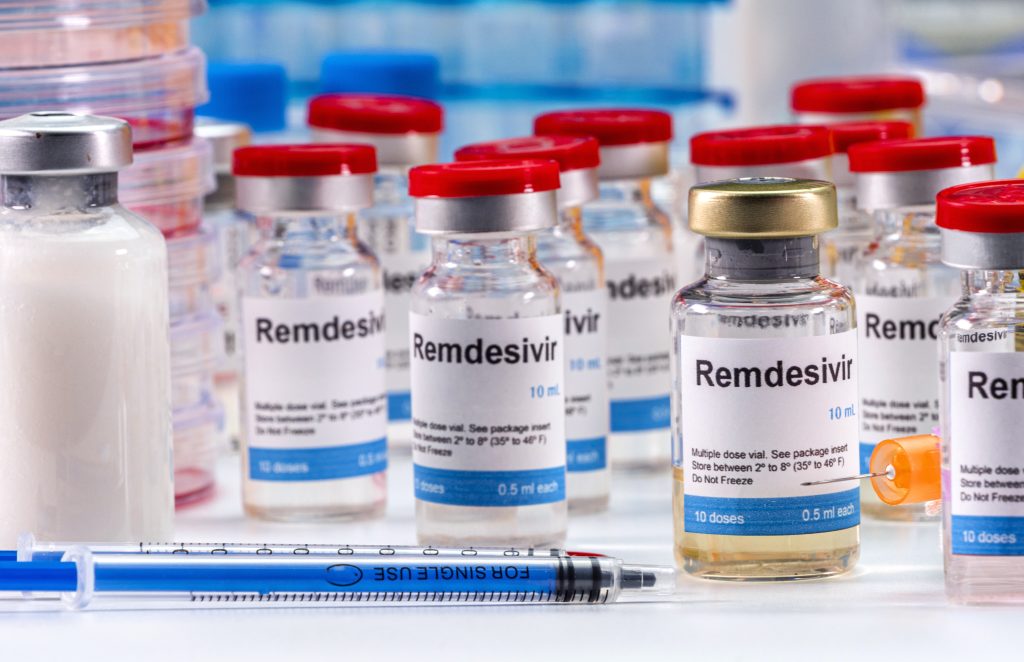
Remdesivir (RDV) is the only Food and Drug Administration approved drug available so far for the treatment of COVID-19. Nucleosides are structural subunits of nucleic acids, and nucleoside analogues resemble naturally occurring nucleosides to form an important class of antiviral agents. RDV is one example of an antiviral compound that was designed to function as a nucleotide analogue, and it can be phosphorylated to closely resemble adenosine triphosphate (ATP), the ubiquitous molecule that carries energy within cells.
RDV was originally developed as an antiviral treatment for the Ebola virus and infections by other coronaviruses, such as Middle East respiratory syndrome and the severe acute respiratory syndrome coronaviruses known as MERS-CoV and SARS-CoV. Critically, RDV works by interfering with viral genome replication and directly competing with substrates of the viral RdRp. But how is this achieved?
Dr Jin Yu from the University of California, Irvine, and her colleagues conducted an elegant computational study to better understand how the nucleotide analogue drug remdesivir (RDV-TP) binds and inserts to the SARS-CoV-2 RdRp active site, in competition with the natural nucleotide substrate ATP.
In order to achieve this, the team first constructed atomic structural models, based on the high-resolution structures determined recently for SARS-CoV-2 RdRp. The study, published in September 2021 in Molecular Systems Design & Engineering by the Royal Society of Chemistry (and featured on the back cover of the issue), shows that RDV is first metabolised into its active form, an adenosine analogue, to interfere with SARS-Cov-2 RdRp. The researchers showed that once in the active site, RDV forms base stacking with RNA template nucleotide and then binds tightly to the enzyme, so that it can be incorporated into the synthesising RNA chain and later on, hinder the RNA synthesis functions of the SARS-CoV-2 RdRp, effectively halting the mechanism of viral replication.
The Advantages of In Silico Investigations Over In Vitro Enzyme Assays
In their 2021 study, Dr Yu and her team simulated the insertion of triphosphate from RDV-TP into the SARS-CoV-2 RdRp active site. The team employed a high-performance computing approach that enabled them to develop a computational microscope to explore the active site of the RNA polymerase. The technique allowed Dr Yu and her team to reveal how natural nucleotides and nucleotide analogue drug candidates incorporate into the RNA chain during viral replication.
The rationale for simulating the nucleotide analogue binding and insertion, rather than monitoring the activity of RdRp in vitro, in the presence of RDV, is that the synthesis of the active nucleotide analogue compound requires considerable time and cannot zoom into molecular dynamics details, to some extent hindering rapid experimental testing. In silico modelling and simulation make it comparatively straightforward to provide insights on the structural and energetics details of the RdRp directed nucleotide incorporation in viral RNA synthesis. Combined with experimental studies, the computational investigations are expected to reveal the physical mechanisms of viral RdRp function, providing the basis for future collaborative efforts on developing drug therapeutics for the treatment of COVID-19.
During the RNA chain elongation process of replication or transcription, a nucleotide binds to the active site of the polymerase (or nearby), subject to initial screening by selectivity of the polymerase and then by collaborative proofreading mechanisms. RdRp recruits ribonucleotides one at a time to the active site and adds the nucleotide to the growing RNA chain. Dr Yu and her colleagues highlighted a complex network of subtle local interactions of amino acids that facilitate the binding and insertion of RDV onto the active site of the viral RdRp. RDV-TP, the nucleotide analogue, disguises itself as a similar ATP substrate for the synthesis of RNA, successfully evading the initial screening, and later, the proofreading step in the RdRp active site.
In fact, the researchers even suggest that the binding and insertion of RDV-TP to the active site of SARS-CoV-2 RdRp can be favoured over that of the natural substrate ATP. The team showed that the RDV-TP initial binding is favoured by forming base stacking with the template RNA nucleotide, and then the RDV-TP insertion is further facilitated and stabilised by hydrogen bonding and salt-bridge interactions between the nucleotide analogue and specific residues around the RdRp active site. Following the successful RDV-TP insertion and incorporation to the viral RNA chain, several nucleotides can be additionally added to the growing RNA chain until termination happens. This is consistent with previous studies that showed that RDV inhibits the Ebola virus and MERS coronavirus via a delayed chain termination mechanism.
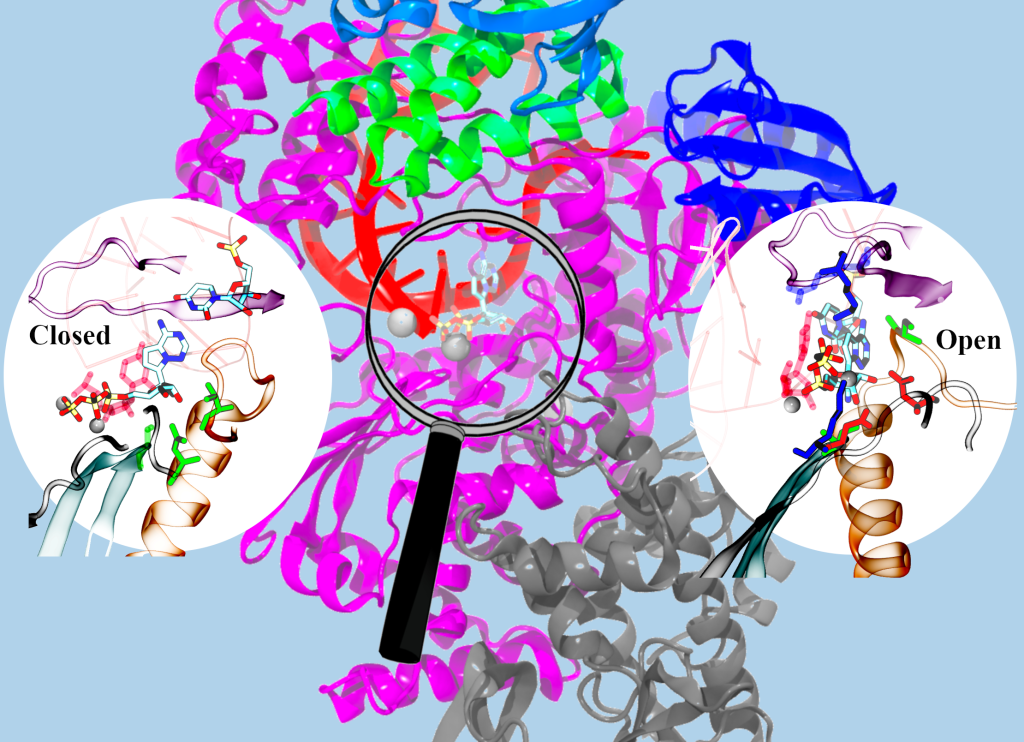
Reproduced with kind permission from the Royal Society of Chemistry
Future Developments
The focus of Dr Yu’s research is aimed at better understanding the biophysics and biochemistry of living systems obtained through a wide spectrum of molecular modelling and simulation techniques. Dr Yu and her team have access to the world’s most powerful high-performance computing resources, the Summit supercomputer from the Oak Ridge National Lab Leadership Computing Facility, that supports COVID-19 research through the COVID-19 High Performance Computing Consortium. This consortium brings together industry, academic and federal agency experts to conduct extensive research in areas like bioinformatics, epidemiology, and molecular modelling to develop strategies to address the threat posed by COVID-19.
The ongoing interdisciplinary studies at the Yu laboratory offer high promise in the development of effective antivirals for the treatment of COVID-19 as well as other diseases. The team’s planned in silico studies will further clarify the physical mechanism of action of RDV and similar analogues and pave the way for the development of additional antiviral compounds and inhibitors of SARS-CoV-2 RdRp. The team also plans to look further into the RNA synthesis mechanisms in SARS-CoV-2, probing for potential drug resistance of the fast-evolving RNA virus.
SHARE
DOWNLOAD E-BOOK
REFERENCE
https://doi.org/10.33548/SCIENTIA775
MEET THE RESEARCHERS

Dr Jin Yu
Department of Physics & Astronomy
University of California, Irvine
California, CA
USA
Dr Jin Yu is an Assistant Professor at the Department of Physics & Astronomy at the University of California (UC), Irvine. Dr Yu obtained her PhD in theoretical and computational biophysics in 2007 from the Department of Physics at the University of Illinois at Urbana-Champaign, under the supervision of Professor Klaus Schulten. After completing her postgraduate studies, Dr Yu received the prestigious UC Berkeley Chancellor’s postdoctoral fellowship. At UC Berkeley, she conducted research on the physical and mathematical modelling of biomolecular machines, such as the viral phi29 DNA packaging motor, with Prof George Oster and Prof Carlos Bustamante. In 2012, she joined the Beijing Computational Science Research Centre as a Principal Investigator, working on the modelling and simulation of viral T7 RNA polymerase transcription and fidelity control. In 2019 Dr Yu returned to the USA, joining the Department of Physics & Astronomy at UC Irvine, where she continues her interdisciplinary studies on computational biophysics. She currently holds a joint appointment with the Department of Chemistry, and an affiliation to the NSF-Simons Center of Multiscale Cell Fate Research at UC Irvine.
CONTACT
E: jin.yu@uci.edu
W: https://www.physics.uci.edu/jin-yu

Moises Romero
Department of Chemistry
University of California, Irvine
California, CA
USA
Moises Romero is a fifth-year graduate student in the Department of Chemistry at UC Irvine. He is the lead author of the current work and has conducted significant modelling and simulation of the SARS-CoV-2 RdRp system. In addition to research, Moises is also heavily involved in diversity, equity and inclusion work, and serves on the board of the Society for Advancement of Chicanos/Hispanics & Native Americans in Science chapter.
CONTACT
E: moiseser@uci.edu
W: https://www.chem.uci.edu/students/graduate-student-160
FUNDING
The current work is supported by the National Science Foundation (Award #2028935)
FURTHER READING
ME Romero, C Long, D La Rocco, et al., Probing remdesivir nucleotide analogue insertion to SARS-CoV-2 RNA dependent RNA polymerase in viral replication, Molecular Systems Design and Engineering, 2021, 6, 888–902.
C Long, ME Romero, D La Rocco, J Yu, Dissecting nucleotide selectivity in viral RNA polymerases, Computational and Structural Biotechnology Journal, 2021, 19, 3339–3348.

REPUBLISH OUR ARTICLES
We encourage all formats of sharing and republishing of our articles. Whether you want to host on your website, publication or blog, we welcome this. Find out more
Creative Commons Licence (CC BY 4.0)
This work is licensed under a Creative Commons Attribution 4.0 International License. 
What does this mean?
Share: You can copy and redistribute the material in any medium or format
Adapt: You can change, and build upon the material for any purpose, even commercially.
Credit: You must give appropriate credit, provide a link to the license, and indicate if changes were made.
SUBSCRIBE NOW
Follow Us
MORE ARTICLES YOU MAY LIKE
Dr Hatim Hassan | Proteins identified in gut bacteria that reduce oxalate levels
New research has identified proteins from gut bacteria, called Sel1-like proteins, that have the potential to help the body get rid of excess oxalate, an organic substance linked to kidney stones, kidney disease, and other health problems. Sel1-like proteins help the cell in assembling large molecular complexes important for cell function. Dr Hatim Hassan from the Division of Nephrology and Hypertension, Mayo Clinic, Rochester, Minnesota, United States, is part of a team of scientists researching whether these proteins and their derived peptides could reduce blood and urinary oxalate levels to prevent and/ or treat hyperoxalemia (high blood oxalate), hyperoxaluria (high urine oxalate) and related disorders (including kidney stones).
Dr Norio Mitsuhashi | Measuring Respiratory Motion to Improve Precision in Lung Radiation Therapy
Dr Norio Mitsuhashi, former Professor of the Department of Radiation Oncology at Tokyo Women’s Medical University, leads revolutionary clinical research into optimising stereotactic body radiation therapy for lung cancer. Dr Mitsuhashi and his colleagues examine whether routinely available patient and tumour characteristics can predict respiratory tumour motion, a critical source of uncertainty in high precision radiotherapy. Their findings suggest that respiratory motion cannot be reliably inferred, and must instead be measured directly in every patient.
Professor Terry C. Hrubec | Clean is good – but is too clean better?
Quaternary ammonium compounds are a large class of compounds used as disinfectants in hospitals, restaurants, healthcare and animal care facilities, and are popular as household cleaners. With disease outbreaks increasing our fears about infections, the use of disinfectants has skyrocketed in recent years. Understandably, we all want to feel safe. However, as Professor Terry Hrubec from the Department of Biomedical Sciences of E. Via College of Osteopathic Medicine discovered, such products may be causing more harm than good.
Professor Abraham P. Lee | Delivering Cancer Immunotherapy with Acoustic-Electric Precision, AESOP’s Fact not Fable
Chimeric Antigen Receptor (CAR) T-cell therapy offers life-saving potential, particularly against blood cancers, but severe side effects such as cytokine release syndrome (CRS) limit its safety. These toxicities are linked to uncontrolled CAR expression levels on the T-cell surface. Led by Professor Abraham P. Lee, researchers at the University of California, Irvine, have developed an advanced microfluidic system, called the Acoustic-Electric Shear Orbiting Poration (AESOP) platform, to precisely control the dose of genetic material delivered into primary T cells. This innovation promises safer, more homogeneous, and highly effective cellular immunotherapies.